Large-scale neurophysiology and single-cell profiling in human neuroscience | Nature
Excerpt
Advances in large-scale single-unit human neurophysiology, single-cell RNA sequencing, spatial transcriptomics and long-term ex vivo tissue culture of surgically resected human brain tissue have provided an unprecedented opportunity to study human neuroscience. In this Perspective, we describe the development of these paradigms, including Neuropixels and recent brain-cell atlas efforts, and discuss how their convergence will further investigations into the cellular underpinnings of network-level activity in the human brain. Specifically, we introduce a workflow in which functionally mapped samples of human brain tissue resected during awake brain surgery can be cultured ex vivo for multi-modal cellular and functional profiling. We then explore how advances in human neuroscience will affect clinical practice, and conclude by discussing societal and ethical implications to consider. Potential findings from the field of human neuroscience will be vast, ranging from insights into human neurodiversity and evolution to providing cell-type-specific access to study and manipulate diseased circuits in pathology. This Perspective aims to provide a unifying framework for the field of human neuroscience as we welcome an exciting era for understanding the functional cytoarchitecture of the human brain. This Perspective considers the implications of advances in human physiology, single-cell and spatial transcriptomics and long-term culture of resected human brain tissue for the study of network-level activity in human neuroscience.
- Perspective
- Published: 19 June 2024
Nature volume 630, pages 587–595 (2024)Cite this article
-
7198 Accesses
-
20 Altmetric
Abstract
Advances in large-scale single-unit human neurophysiology, single-cell RNA sequencing, spatial transcriptomics and long-term ex vivo tissue culture of surgically resected human brain tissue have provided an unprecedented opportunity to study human neuroscience. In this Perspective, we describe the development of these paradigms, including Neuropixels and recent brain-cell atlas efforts, and discuss how their convergence will further investigations into the cellular underpinnings of network-level activity in the human brain. Specifically, we introduce a workflow in which functionally mapped samples of human brain tissue resected during awake brain surgery can be cultured ex vivo for multi-modal cellular and functional profiling. We then explore how advances in human neuroscience will affect clinical practice, and conclude by discussing societal and ethical implications to consider. Potential findings from the field of human neuroscience will be vast, ranging from insights into human neurodiversity and evolution to providing cell-type-specific access to study and manipulate diseased circuits in pathology. This Perspective aims to provide a unifying framework for the field of human neuroscience as we welcome an exciting era for understanding the functional cytoarchitecture of the human brain.
Similar content being viewed by others
Main
Modern neuroscience began with Paul Broca’s discovery of language cortex, which was based on observations that lesions in the human brain disrupt normal cognitive functions. Subsequently, Wilder Penfield went on to map the functional areas of the human cerebral cortex using intraoperative stimulation to localize language, memory and movements. Subsequent mechanistic studies aiming to understand brain function beyond localization have primarily been advanced using animal models. Although these studies reveal fundamental principles about the formation and function of neural circuits, this has left a gap in our understanding of regulators in higher functional domains such as language and higher sensory processing. Uncovering the genetic, cellular and molecular bases of cognitive specializations observed in humans (Box 1) has prompted renewed interest in studying the human brain. However, studies of the human brain are challenging, owing to the lack of scalable tools to manipulate cell behaviour or gene expression and confounding variables of individuality.
In this perspective, we highlight emerging efforts to advance human neuroscience through our personal experiences as a collaborative team focused on cellular-level function and structure, including neurosurgeons (E.F.C. and A.T.L.), a neurologist (M.F.P.) and basic neuroscientists (T.J.N. and A.T.L.). We believe that the convergence of neuroscience research techniques that enable mapping of the human brain at a large scale and with cellular resolution has arrived for the fields of human neurophysiology and anatomy to herald the burgeoning field of human neuroscience. Specifically, we describe how parallel breakthroughs in the surgical operating room, including the ability to perform large-scale recordings of neural activity in single neurons in awake behaving participants, coupled with long-term culture methods for resected brain tissue, open up an unprecedented opportunity to study the entire spectrum of human neuroscience, from network, to microcircuit, to cell types and their molecular features. Given our collective expertise, we focus not specifically on advances in cognitive and systems neuroscience, but rather on how we can leverage human tissue to study cellular function and genetic diversity. Notably, as we make strides in understanding the genetic, cellular and molecular bases of human cognitive specializations, we face important questions about societal implications and ethics related to human neuroscience research. We describe paradigms that may enable the scientific community to proactively engage in broader societal domains, including the concepts of informed consent and data sharing—especially in a field in which patients are experimental subjects, direct benefit from the research is currently limited, and crucially, the ramifications of future advances in machine learning and artificial intelligence (AI) analyses remain unknown.
Large-scale neurophysiology of human behaviour
Non-invasive imaging technologies such as functional magnetic resonance imaging (fMRI) have provided important insights into neural dynamics across the entire human brain, but are substantially limited in resolution. Technologies that can directly record neural activity from the human brain have improved substantially over the past decade, increasing in both scale and spatiotemporal resolution.
Historically, traditional tungsten microelectrodes used in intraoperative recordings measured activity from a few neurons per electrode. Despite their limitation of scale, these devices have high spatiotemporal resolution, which has enabled direct measurements of activity from single neurons, for example in the basal ganglia for placement of deep brain stimulation electrodes in Parkinson’s disease. Recordings from stereotactically guided depth electrodes in stereoelectroencephalography and placement of craniotomy-based surface grid electrocorticography electrodes (ECOG) in epilepsy surgery are able to capture large neuronal networks, but lack the spatiotemporal resolution necessary to detect single action potential spikes. Instead, these latter modalities capture an aggregate activity of inputs to local neuronal networks via local field potentials (LFPs). Thus what has been lacking in the field of human neurophysiology is a means of capturing single units at large scale.
Understanding single-unit activity is critical for a fuller understanding of network function. First, inferences of neuronal spiking activity from LFPs alone are indirect1 and do not address fundamental questions of functional organization, circuit connectivity or information processing2. Indeed, how oscillations (as detected by LFPs) contribute to cell-type-specific neuronal firing patterns and vice versa is an area of active investigation in model systems3,4,5. In rodents, specific inhibitory neuron subtypes respond preferentially to various aspects of sensory discrimination tasks, and different oscillation frequencies are associated with a range of disparate cognitive functions6,7,8,9,10,11 (such as theta in working memory and beta in rule application). For example, inhibition of prefrontal parvalbumin interneurons in mice reduces gamma oscillations and negatively affects rule-shifting tasks4. How these findings from model systems translate to human neuroscience is an open question, notwithstanding the cognitive functions, such as language, that cannot be readily studied in model systems. From a neuroengineering perspective, multiscale decoding algorithms that incorporate both LFP and individual spikes outperform single-scale decoding using either modality alone in brain–machine interface simulations12.
Previous approaches to obtain higher yields of single-unit recordings include electrode designs such as microwire bundles that exit depth electrode tips (Behnke–Fried electrode) and the microelectrode Utah array, which permits action potential recordings from dozens of single units. Indeed, some groups have been able to leverage these techniques to collect thousands of single-neuron activities, although these studies require large cohorts and hundreds of separate individual probes13. These methods, however, are limited and have not advanced substantially since their introduction several decades ago. For electrode technologies to be applied in human neuroscience, they must address several critical issues, including: (1) spatial resolution (that is, action potentials of single cells); (2) temporal resolution on the order of microseconds; (3) brain coverage (local ensemble or cortical column, or across disparate areas); (4) robustness (ability to record for minutes intraoperatively, versus chronically for months and years); and (5) safety (biocompatibility and lack of injury from insertion).
A recent fundamental advance is the Neuropixels probe, which features 1,000 electrodes on a narrow shank, with the potential to record from hundreds of neurons with a single probe insertion. Neuropixels probes have transformed electrophysiology research in rodents and non-human primates14. We recently demonstrated their feasibility for simultaneous large-scale data collection from single neurons in human intraoperative recordings after overcoming challenges such as probe fractures, insertion difficulties, electrical noise and brain movement-related artifacts15,16,17. The vertical probe orientation allows hundreds of cellular recordings within a cortical column, across all cortical layers. Recordings appear robust within seconds of insertion and thereby overcome the challenges of previous single-unit recordings with low yield. Of note, the data acquisition system is turnkey and is constantly improved by a community of active researchers. Recent advances in Neuropixels have overcome existing limitations in non-human primate neurophysiology, such as large-scale sulcal recordings or simultaneous widespread recordings from multiple interconnected brain regions such as primary motor, supplemental motor area and basal ganglia for studying motor planning and control<a data-track=“click” data-track-action=“reference anchor” data-track-label=“link” data-test=“citation-ref” aria-label=“Reference 18” title=“Trautmann, E. M. et al. Large-scale high-density brain-wide neural recording in nonhuman primates. Preprint at bioRxiv https://doi.org/10.1101/2023.02.01.526664
(2023)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR18" id="ref-link-section-d35850473e641">18</a></sup>.
Using Neuropixels, we have addressed questions about stimulus feature encoding in speech that have been difficult to address using LFP recordings alone. Because Neuropixels probes sample spiking activity through all cortical layers, we now have the capability to study how neuronal activity is processed as it flows through cortical layers within an individual cortical column. Although model systems have begun to address questions of cortical columnar processing, many of these techniques require calcium dyes, viral transduction of calcium or voltage indicators, two-photon imaging or implantation of large lenses, none of which has the temporal resolution or safety feasibility of Neuropixels for human research19. By integrating population-level LFP data with the spiking activity of individual neurons and information processing through cortical layers, Neuropixels recordings afford us opportunities to study the cellular mechanisms of how auditory circuits encode the multitude of stimulus features in human language within cortical columns and across cortical layers17,20. Future work will prioritize scaling Neuropixels probes so that recordings can be carried out in parallel from multiple simultaneous brain regions, sampling tens of thousands of single neurons, and ideally being fully compatible with chronic recordings. One limitation of the current Neuropixels technology is that an individual probe can collect data from only a single cortical area. To facilitate larger cortical coverage, emerging single-neuron recording techniques could be distributed over a wider range of the cerebral cortex. Another complementary approach includes high-density micro-electrocorticography20,21, which focuses on recordings from the cortical surface using new materials and fabrication processes that are common to advanced micro-electromechanical systems devices, enabling electrode arrays to be fabricated using chip printing lithography on highly flexible and conformal polymer substrates. Such approaches have the advantage of covering substantial cortical areas while maintaining the safety of avoiding cortical surface penetration. Recent demonstrations include the ability to record action potentials, as well as very focal cortical events like high-frequency oscillations (HFOs), on the scale of ten of micrometers in resolution22. Together, surface ECOG and penetrating laminar multi-electrode arrays such as Neuropixels will enable unprecedented three-dimensional measurements from cortical and subcortical structures to increase understanding of how distributed neuronal activity gives rise to patterned activity and human behaviours23.
An atlas of cell types and circuits
Relating the activity of a single neuron measured in the context of an in vivo circuit or network to the molecular identity of that neuron represents a fundamental and unresolved challenge in neuroscience. A deeper appreciation for the cellular building blocks of a neuronal circuit will not only provide more mechanistic understanding of how different types of information are processed in the human brain, but also potentially provide opportunities to modulate physiology or the transcriptome in diseases with cell-type-specific treatments. Approaches such as a priori genetic labelling of molecular cell types followed by in vivo activity recordings have provided some of the first proof-of-concept applications, but face challenges in application to non-model organisms, including humans24.
Cells in the brain have been classically categorized into types on the basis of morphology, anatomical position and intrinsic physiological properties, such as action potentials, resting membrane properties and various channel properties. More recently, single-cell transcriptomics and epigenomics have enabled researchers to capture a more complete spectrum of cellular properties thought to define cell types (Fig. 1). We initiated these efforts in brain tissue by profiling dozens of cells25, then went on to report hundreds and thousands of cells26,27. Breakthroughs in technology for capturing single cells using microfluidics have opened up a practical path towards census-scale efforts in neuroscience28, many of which are currently ongoing. Recently, a complete census of the mouse brain has been completed, reporting that the adult mouse brain contains approximately 5,000 cell types29,30. An initial draft atlas of the human brain revealed over 400 clusters and 3,000 subclusters of cell types, with many showing regional diversity in neurons and macroglia unique to their respective forebrain, midbrain, and hindbrain regions31. How this complexity compares between the mouse and human brain human brain is likely to be resolved over the next few years. Advances in technology that enable microfluidics-free cell capture32 will further accelerate these efforts by enabling routine capture of tens to hundreds of millions of cells in a given study.
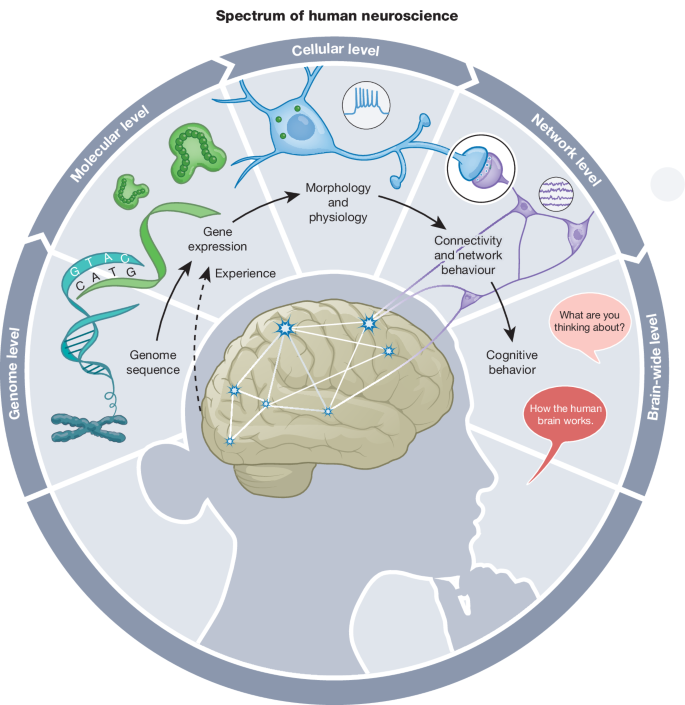
Fig. 1: The central dogma of neuroscience.
Cells in the human brain share the same genomic information but develop distinct patterns of gene expression. In turn, gene-expression profiles correlate with and, in many cases, regulate the abundance of proteins that build the cells and determine their distinct properties, such as morphology and physiology. Ensembles of neuronal cells give rise to synaptically coupled neuronal networks that generate network-level properties. Coordinated activities of neurons in neural circuits give rise to motor output and cognition. Importantly, neuronal activity feeds back to control gene expression through activity-dependent molecular and cellular changes. Figure originally drawn by Noel Sirivansanti.
Ongoing efforts seek to characterize the diversity of cell types in the human brain using single-cell or single-nucleus transcriptomics33, chromatin state profiling and epigenomics34,35,36. New and emerging efforts are beginning to extend the spectrum of methodologies to incorporate measurements of lipids, metabolites and proteins37, opening up new opportunities for deep molecular characterization of cell types in the human brain. It is important to recognize that many features of cell identity, such as the subcellular transcriptome, are not captured in those studies owing to the limitations of current technologies38,39. We must also embrace the fact that a consensus definition of a cell type is currently lacking40, but even if the question of the absolute number of cell types in the brain cannot be fully addressed, we can be successful in defining their transcriptomic marker genes. The effects of census-scale transcriptomic studies have already led to the development of technologies that enable us to visualize transcriptomic cell types in tissue38,41,42,43 and begin to precisely localize them within a common coordinate framework44.
Linking cellular function and networks
Molecular cell states have primarily been assayed using post mortem human tissue specimens. However, the time needed to collect and freeze post mortem tissue may result in significant changes in gene expression<a data-track=“click” data-track-action=“reference anchor” data-track-label=“link” data-test=“citation-ref” aria-label=“Reference 45” title=“Liharska, L. E. et al. A study of gene expression in the living human brain. Preprint at medRxiv https://doi.org/10.1101/2023.04.21.23288916
(2023)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR45" id="ref-link-section-d35850473e794">45</a></sup>. Furthermore, studies of post mortem tissue do not capture the functional properties of cells, many of which could be fundamental to cell-type definition (Fig. [1](https://www.nature.com/articles/s41586-024-07405-0#Fig1)). The extent to which transcriptomic cell types contribute to network dynamics is an open question that cannot be fully addressed even in model organisms<sup><a data-track="click" data-track-action="reference anchor" data-track-label="link" data-test="citation-ref" aria-label="Reference 24" title="Bugeon, S. et al. A transcriptomic axis predicts state modulation of cortical interneurons. Nature 607, 330–338 (2022)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR24" id="ref-link-section-d35850473e801">24</a>,<a data-track="click" data-track-action="reference anchor" data-track-label="link" data-test="citation-ref" aria-label="Reference 46" title="Schneider, A. et al. Transcriptomic cell type structures in vivo neuronal activity across multiple timescales. Cell Rep. 42, 112318 (2023)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR46" id="ref-link-section-d35850473e804">46</a></sup>. Thus, the advent of approaches that allow neurosurgical specimens to be maintained viable in culture for multiple days offers an alternative opportunity to study cell types within a functional microcircuit<sup><a data-track="click" data-track-action="reference anchor" data-track-label="link" data-test="citation-ref" aria-label="Reference 24" title="Bugeon, S. et al. A transcriptomic axis predicts state modulation of cortical interneurons. Nature 607, 330–338 (2022)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR24" id="ref-link-section-d35850473e808">24</a>,<a data-track="click" data-track-action="reference anchor" data-track-label="link" data-test="citation-ref" aria-label="Reference 46" title="Schneider, A. et al. Transcriptomic cell type structures in vivo neuronal activity across multiple timescales. Cell Rep. 42, 112318 (2023)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR46" id="ref-link-section-d35850473e811">46</a>,<a data-track="click" data-track-action="reference anchor" data-track-label="link" data-test="citation-ref" aria-label="Reference 47" title="Ting, J. T. et al. A robust ex vivo experimental platform for molecular-genetic dissection of adult human neocortical cell types and circuits. Sci. Rep. 8, 8407 (2018)." href="https://www.nature.com/articles/s41586-024-07405-0#ref-CR47" id="ref-link-section-d35850473e814">47</a></sup> (Fig. [2](https://www.nature.com/articles/s41586-024-07405-0#Fig2)).
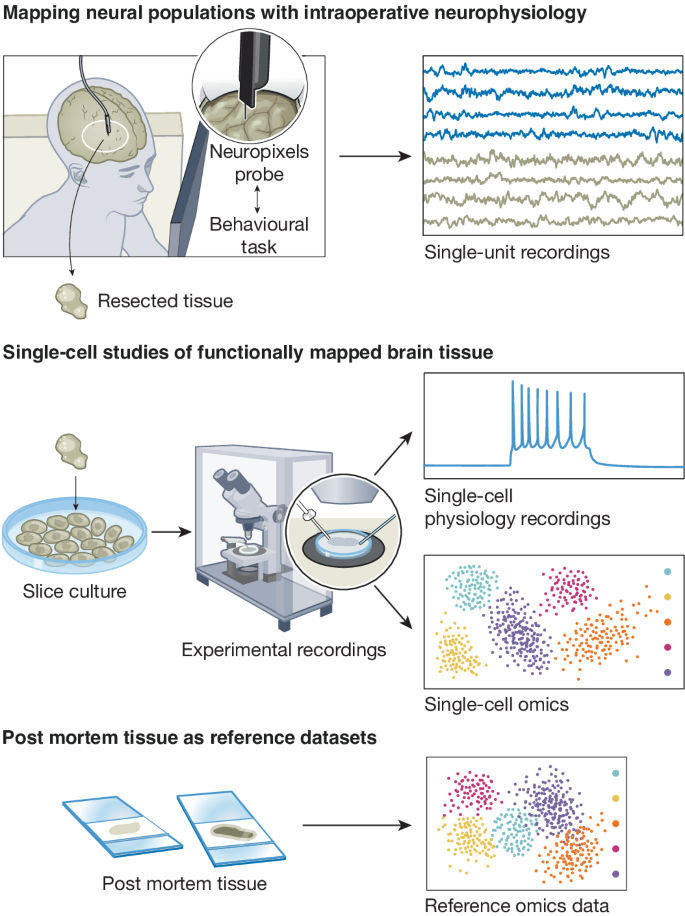
Fig. 2: Central approaches to advance the study of human brain.
Intraoperative recordings of neural activity while patients are doing cognitive tasks can be conducted ethically and with patient consent to gain insight into neural correlates of human behaviour (top). Tissue resected from the brain can be maintained in ex vivo culture condition to facilitate single-cell electrophysiology and molecular profiling (middle). In parallel, post mortem tissue specimens provide generalizable insights across brain regions and broader patient population (bottom). Figure originally drawn by Noel Sirivansanti.
Human brain tissue resected from surgery has already begun to reveal new insights into how human neurons process information by leveraging human-specific neurophysiology such as unique dendritic compartmentalization and computation48,49,50,51. As many types of surgery for epilepsy require patients to be awake, ex vivo approaches also afford a rare opportunity to investigate the genetic and cellular composition of tissue that has been functionally mapped as the patient undergoes cognitive testing. For example, transcriptomic studies of resected tissue functionally mapped with episodic memory tasks have linked specific gene-expression networks with oscillatory signatures of memory formation across multiple frequency bands52. Indeed, the ability of the ex vivo approach to dissect the cellular architecture of brain tissue whose networks have been functionally mapped by Neuropixels or ECOG during human cognitive tasks (that is, language) is a key distinction between neuroscience in humans versus other model systems.
Studying neurosurgically resected tissue in culture also provides a window to study subtype-specific synaptic connectivity maps, which will be critical in linking cell types to circuit and network function. Although advances in electron microscopy will soon enable whole-mouse brain connectome studies53, achieving the scale necessary for the human brain will be more challenging. Moreover, strategies to visualize molecular cell types in electron microscopy workflows are less well developed. Virally introducing synaptically connected molecular barcodes into ex vivo slices and then profiling their connectivity with next-generation single-cell sequencing technologies may enable us to overcome the limitations of cell-type identities and scale seen in electron microscopy connectome strategies54. Although ex vivo virally mediated barcode-tracing strategies inherently narrow study to the local microcircuit within each slice, the integration of cellular identity and single-cell transcriptomics with synaptic connectivity maps may highlight intrinsic differences in endogenous network activity motifs between healthy and disease states55,56,57.
To take this further, we need to find efficient ways to relate in vivo cellular activity to cell-type identity, such as developing approaches that can register the location of a cell during in vivo recording and determine its identity ex vivo. By preserving the locations of cells from in vivo recordings to their ex vivo positions, large multidimensional datasets that incorporate neurophysiology with single-cell sequencing techniques (transcriptomics, chromatin state profiling and epigenomics) can be built. This can be accomplished to some degree using extracellular electrodes58. An alternative approach may be to leverage in silico models, which provide biorealistic approximations of brain function. Success of biorealistic in silico models trained with data from humans will provide a representation, and predictions from such models can be tested for validation59,60.
Several issues with the ex vivo tissue culture approach need to be considered, including: (1) the generalizability of findings from tissue collected in epilepsy surgeries; (2) the additional technical variation of cell health in tissue culture; and (3) the constraints of studying a microcircuit in isolation. As the majority of tissue used in these studies is derived from patients with epilepsy, an important caveat is that conclusions from these studies may not routinely be generalizable to the broader population. However, this limitation does not necessarily discount the utility of insights derived from this patient cohort. At our institution, a substantial proportion of our patients with epilepsy undergo focused temporal lobectomies, in which the primary epileptogenic pathology is in subcortical structures such as the hippocampus and amygdala, and not necessarily in the cortex. Moreover, because experimental conditions are compared across patients with epilepsy rather than control cohorts, other control methodologies can be used, such as matched post mortem gene-expression samples from people with epilepsy and control individuals52. In fact, the heterogeneity of patients with epilepsy may itself pose a unique research opportunity, as this population often harbours co-morbidities such as depression, anxiety or autism, which may also be of interest. Furthermore, multiple non-neuronal tissue types, such as endothelial cells of the cerebrovascular system, can also be studied in tissue from patients with epilepsy, and these findings may be more broadly generalizable61. Similar considerations of generalizability would also apply for tissue collected during resections of deep brain tumours. Overall, ex vivo human neuroscience approaches can be used to stimulate growth in the field and promote hypotheses that can be supported with other techniques including gene-association, exome-sequencing or proteomic studies. Regarding technical variations introduced by tissue culture, neurons from cultured brain slices have been demonstrated to have stable electrophysiological properties over several weeks and are similar to acute slices, as evidenced by patch-clamp electrophysiology47,62,63. Finally, although the ex vivo approach results in microcircuits lacking their native in vivo inputs, we can still study questions of subtype composition, synaptic connectivity, cell-intrinsic physiology and endogenous network motifs, which together can provide mechanistic insights into the larger intact network.
Finally, human ex vivo tissue culture models can provide a number of unique opportunities to advance therapeutic opportunities. First, defining cellular states directly in diseased tissue can reveal unexpected cellular, genetic and intercellular phenotypes that could reveal unexpected disease transformations or changes that correlate with disease progression, and might inform cross-model studies of animal and in vitro systems61. The process of developing and validating new vectors for gene therapy, such as cell-type-specific enhancer discovery64 or adeno-associated virus (AAV) capsid engineering by directed evolution, could be accelerated by incorporating human tissue-based experiments, thereby lowering the risks of such tools not translating from mouse or monkey to human65.
Human neuroscience and clinical practice
So far, we have described approaches to advance human neuroscience at the cellular and network levels, and how to connect those dimensions to typical brain function. Although these advances will yield exciting insights into what makes us human, they also highlight the unmet need to develop strategies to precisely modulate the neural network at the gene, cellular or circuit level for the benefit of patients with neurological or psychiatric conditions. This journey starts with the need to systematically define cellular function and dysfunction in the context of disease and, with rapid gains in basic science knowledge, to develop new precision tools for cell- or circuit-specific treatments.
Genetic mapping and precision medicine
Genomic, transcriptomic and proteomic studies have defined a starting point for understanding the neuropathological mechanisms of many neurological and psychiatric conditions. Gene-association and exome-sequencing studies have begun to identify the genetic risk architecture of diseases such as motor neuron disease66, neurodegenerative diseases67,68, epilepsy69,70, autism spectrum and neurodevelopmental disorders71,72, and schizophrenia73,74. Rapid advances in sequencing technologies have enabled the application of genetic diagnosis75 to improve diagnostic capabilities. The identification of variants in risk genes for complex conditions such as autism spectrum disorder and epilepsy have been increasingly incorporated into clinical work-ups76,77 and can provide guidance when symptoms point to a suspected diagnosis.
However, genomic variants identified at the plasma level explain a small fraction of cases of many neurological and psychiatric conditions78. The ability to access human brain tissue, such as via neurosurgical resections, has revealed a unique set of mutations that emerge outside the germline, leading to a growing appreciation for somatic mosaicism in the role of typical brain development and in neurological disease79,80,81,82. Mutations in this scenario emerge later in development, after the zygote state, leading to organ tissues that are mosaic, with only a subset of cells containing the mutation. Such somatic mutations have been observed specifically in tissue obtained from patients undergoing neurosurgery, such as those with focal epilepsy who have undergone resection to remove a lesion, as in cortical dysplasia or hemimegalencephaly83,84. This has led to the identification of new disease risk genes for focal cortical dysplasias and hemimegalencephaly, including AKT3 (ref. 85) and DEPDC5 (ref. 86), and a converging role of the mTOR pathway in neurodevelopment-associated epilepsy87. These findings would not have been possible without the interrogation of human neurosurgical material.
Gain- or loss-of-function of genes identified in these studies increase the risk for a particular condition, and this information can guide pharmaceutical development, or even repurposing of old drugs. For example, Dravet syndrome is caused by loss of function mutations in SCN1A, which encodes a sodium channel. Identification of the causal gene has led to practical ways to improve patient outcomes, including better treatment strategy (avoiding certain seizure medications) and the use of pharmacological agents such as fenfluramine and cannabidiol88.
Furthermore, mutational burden information can be integrated with cell-type-specific gene-expression information profiles to highlight patterns of select cell-type vulnerability to disease-causing mutations. One example of this is the identification of TREM2 as a genetic risk factor for Alzheimer’s disease89. TREM2 is selectively expressed in microglia90, a finding that has sparked substantial efforts to understand the role of the innate immune system in Alzheimer’s disease. Of course, gene expression in any one cell type may not be necessary or sufficient to explain the full spectrum of disease symptoms, but identifying profiles of gene expression may represent an important step towards systematic interrogation of cell-type-specific contributions of patient-specific mutations to disease processes. Such information can also guide efforts to develop improved cellular models that could serve as platforms to interrogate disease mechanisms and serve as preclinical models.
Although the vast majority of single-cell reference atlases of human cell-type diversity do not extensively characterize mRNA isoforms, an important yet underestimated ramification of cell-type-specific atlases is the potential to identify pathogenic and cell-type-specific mRNA isoforms. Antisense oligonucleotides (ASOs) are single-stranded synthetic short molecules that bind to target mRNA and lead to target-specific knockdown. ASOs have been used to treat Duchenne muscular dystrophy91,92 and spinal muscular atrophy93. The biggest limitations of the use of ASOs are ensuring correct sequence matching to target genes (to optimize gene targeting of all isoforms and prevent off-target effects) and delivery, as these molecules are short-lived and cannot cross the blood–brain barrier. Isoform-resolved single-cell transcriptomic atlases could accelerate the identification of potential targets for this emerging class of therapeutic tools.
The ability to integrate the increasing datasets available94,95,96,97 into machine learning-enabled formats98 will enable deep learning approaches to combine the genetic, clinical and environmental factors into a more cohesive framework to create tailored and accurate therapeutic strategies.
Cell-type-specific molecular access
The emergence of tools for cell-type-specific molecular access to monitor, target and manipulate specific cell types in the human brain opens up opportunities to pursue mechanistic and therapeutic studies. Gene therapy offers a direct way to repair or replace a defective gene, an approach that is highly suited for monogenic conditions. AAV vectors represent an approach that has been successfully applied to target nervous system tissue99,100. AAV vectors can infect many cell types, show low immunogenicity and enable long-term expression of introduced transgenes without requiring integration into the host chromosome. One successful example in neurology is spinal muscular atrophy, a devastating neurodegenerative neuromuscular disease101 caused by mutations in SMN1 or SMN2 (ref. 102). The positive results from the initial cohort have led to enthusiasm about the potential for in vivo gene therapy, such as via AAV vectors, and have served as a guide for gene therapy trials in other conditions103. There are currently trials to treat conditions such as Canavan disease (with mutations in ASPA, which encodes aspartoacylase), Batten disease (a lysosomal storage disorder caused by the loss of tripeptidyl peptidase 1) and aromatic-l-amino acid decarboxylase deficiency104,105. However, most AAVs used in current clinical trials have very broad tropism, and as we begin to better understand neurological disease at the cell-type and circuit level, it will become increasingly important to develop cell-type-specific AAVs. Advances in the engineering of AAV capsids and identification of cell-type-specific enhancer elements will enable more precise AAV vector targeting106,107,108 and applicability across species64. This next generation of vectors will enable the delivery of a variety of effector molecules, not limited to causal disease genes, and will be increasingly tailored to cargos such as optogenetic or chemogenetic actuators of neural activity.
Ethics and data in human neuroscience
Human neuroscience research depends critically on patient volunteers. Participation should involve informed consent, which describes both the risks of tissue collection and neural device implantation and details of how the data generated can be used and shared with the research community. This is especially important because the immediate benefit of surgical patients participating as research subjects is currently limited and the risks are potentially severe, in both obvious and unpredictable ways. For tissue collection, it is paramount that individuals understand that participation in tissue collection does not change their surgical plan. Only tissue that is clinically indicated for resection is collected for research. The ethical frameworks for human research with neural devices have been reviewed extensively by the NIH BRAIN (Brain Research Through Advancing Innovative Neurotechnologies) Initiative Neuroethics Working Group and include the analysis of risk, informed consent, and post-trial responsibilities to research participants undergoing implantation of neural devices109. With regards specifically to data sharing, the Working Group states that the social and scientific value of data sharing should be weighed against robust analyses of privacy risks, and that sharing data may be a key post-trial responsibility. In practice, many types of human neurophysiology data are intrinsically difficult to de-identify, particularly the high-dimensional datasets obtained from recording technologies described earlier in this Perspective. However, informed consent should recognize that unforeseen advances in machine learning and AI may risk patient de-identification, particularly as neurophysiology data begin to be superimposed with the individuals’ corresponding transcriptomics, epigenomics and proteomics datasets. The merging and analyses of these complex datasets will require the computing and analytical power of machine learning and AI. In this context, what constitutes informed consent will need to be revisited as we increasingly rely on these technologies, and the degree to which their conclusions arise from a spectrum of ‘white-box’ or ‘black-box’ approaches110 (that is, explainable versus non-explainable models). In the near future, an important discussion for all stakeholders is what safeguards to have in place as AI advances towards artificial general intelligence111. One way to maintain faith in the informed consent process is for the academic research community to make transparent any partnerships between academics and industry, particularly as private industry is an important driver of AI advances.
Recently, the Research Opportunities in Humans Consortium has developed a series of best practices around engaging patients in research to maintain the integrity of clinical care and ensure voluntariness of participation112. However, further discussions are needed, especially around data-sharing practices. Considerations around the privacy and ownership of research data have been quite advanced in other fields such as genomics, where public benefits to datasets involving large numbers of study participants are more clear. Questions remain on the ramifications of broad dissemination of data, who should have access, the public benefits and risks. Answers to those questions are less clearly defined in neuroscience than in genomics.
Sharing research findings based on data derived from study participants is generally viewed as safe, because the raw data are de-identified or because the underlying raw data did not carry information that would pose a possibility of re-identifying the study participants. New data-sharing policies that set FAIR (findability, accessibility, interoperability and reusability) principles at the centre of federally funded research are set to revolutionize biomedical research113, but also prompt a reflection on the risks of data sharing. Even when data are de-identified, with the advent of very large-scale datasets, the possibility of re-identification becomes a point of concern. Nowhere is this more apparent than in the area of genomics, where naturally occurring human genetic variation can be used with some degree of confidence to re-identify an individual114, which is particularly important as commercial providers of genome sequencing create incentives to share genomic material of a large proportion of the population115. To account for the possibility of re-identifying study participants, consent forms now often implement explicit language informing the participants about the potential risk that the data generated as a result of their participation could be used to re-identify them. However, as discussed above, it is difficult to fully predict how future technologies or analytical approaches might open up new avenues to use the data. To limit the risk of potential misuse of such information, researchers can make study data available through controlled access archives, which impose restrictions on how the data can be utilized. At this point, these nuanced considerations have been largely limited to genomic data because of the clear evidence that genetic sequence information is unique to every individual and has implications for clinical management116,117.
In genomics, population-scale studies have demonstrated the importance of sequence-level variation as it relates to identifying individuals with genetic risk who could be targeted for screening and early detection, which in the case of cancer mutations can substantially increase survival118. In neuropsychiatric disease, recent investigations in somatic copy-number variants have detected potential genetic loci for increasing schizophrenia risk119. In terms of affecting medical management, multiple examples exist where information about a patient’s genetic background can lead to a more precise treatment plan, such as changes in clopidogrel metabolism based on CYP2C19 genotype120 and increasing progression-free survival using targeted isocitrate dehydrogenase inhibitors in isocitrate dehydrogenase-mutant glioma121. Thus, genomics information has been transformative thanks to two properties: genetic sequence information unique to every individual, and the implications such information has for disease risk assessment, prevention and treatment strategy. Another revolution poised to take advantage of genomic variant detection includes recent developments in CRISPR therapies122.
Although neuroscience data currently have no clear equivalents of these two properties, ongoing large-scale data-generation efforts will begin to test this assumption. Single-cell genomics datasets are already beginning to uncover molecular states of cells associated with diseases such as Alzheimer’s disease123,124,125. The advent of systematic taxonomies of cell types in the human brain will provide a quantitative structure to systematically compare these cellular phenotypes across disorders126, and the knowledge generated as a result of such comparisons will probably lead to therapeutic interventions that precisely target the underlying cells and circuits. Although the full potential of single-cell atlases has not been fully demonstrated, there are multiple ways in which such data are likely to contribute to addressing treatment needs in neurological disorders, as discussed above.
What is less apparent is how data collected during intraoperative neural recordings should be managed. It is clear that neural activity data carries important information about the underlying disease states, and large-scale datasets collected across hundreds of individuals will probably reveal common patterns of neural function that could provide inroads into personalized therapy. In this scenario, data sharing would ultimately benefit individual patients. However, as transcriptomic and epigenomic information are superimposed onto the physiology datasets, human neuroscience may begin to uncover the genetic and physiological codes that underlie our cognitive functions at the personal and population levels. The research and broader communities will need to be prepared about how to explore these questions, as there is real potential for group-level harm and the use of data to affirm existing biases against certain groups of people. It will require the neuroscience community to engage not only with patients, but also with the general public, about the ramifications for broad dissemination of data, the public benefits and risks of different degrees of data access, and the perspectives of different types of community (such as race, ethnicity and faith) on how to conduct and apply findings from human neuroscience research. At first glance, it is sensible that the sharing of neural activity data would be well served to follow similar principles to genomic information. This approach ensures that all findable, accessible, interoperable and reusable datasets collected from an individual could be removed if requested by the patient. However, patient and community engagement will be just as critical to support a diversification of neuroscience research participants, which is required for an understanding of human brain function that is inclusive of all people.
The success of human neuroscience
Together, by understanding fundamental principles of human brain structure and function, mechanistically interrogating genetic, cellular and neurophysiological properties of neurons and glia, and tdeveloping tools to precisely monitor, target and replace cells of the brain, we will begin to deliver on the ambitious goals of the BRAIN initiative. But it remains to be determined how can we best capitalize on these discoveries and technological advances, what the future of human neuroscience holds now that we are beginning to gain the right tools and mechanisms to study the human brain, and how this should be done responsibly and ethically.
One of the most transformative lessons from the Human Genome Project has been that, in order to enable rapid data and knowledge sharing, the scientific community needs to adopt the use of accessible data-sharing platforms and data browsers, which organize data in a common reference form. Similar efforts are beginning to organize around proposals for cellular taxonomies126, but at the current pace of data production, we predict an even more pressing need to accelerate efforts for adoption of cloud-enabled data repositories127 and tools for integrative analysis and annotation, including large language models. The focus of such analyses must include not only the definition of cell types and states but also phenotypes, as opposed to reference taxonomies, as well as the development of comparable tools for epigenetic datasets that compare chromatin accessibility. Non-discrete, continuous sources of variation related to cellular trajectories that may exist during development or state transition may pose considerable analytical challenges. Responsible data sharing and use policies for genetic variant information must be considered128. There is an emerging but unmet need to develop standards of data sharing in neurophysiology that could serve as a foundation to accelerate secure data platforms for the analysis of human neurophysiological datasets derived from surgically implanted probes, especially at the time when commercial entities begin to consider adoption of such technologies.
The second key issue is that an approach to science will have to fundamentally change from projects driven by single investigators to community-driven efforts, as evidenced by similar collaborative human neuroscience efforts to ours across the USA (for example, Massachusetts General Hospital, Stanford, Baylor, Southwestern and Cedar-Sinai). This will bring several major benefits. Partnerships involving basic science researchers, computational scientists, physician-scientists and physicians will galvanize projects that interrogate questions in human neuroscience across scales and across models. We argue that such partnerships should include engagement efforts with communities of interest, including patient groups and study participants who ultimately consent to share their information, data and tissue information in the trust that such data will be used in a responsible manner and could ultimately lead to a better understanding of neurological conditions and neurodiversity, and to the development of tools to modulate neural functions. We believe that a revolution in biomedical science has already begun, as evidenced by the formation of BRAIN Initiative Network efforts129 to tackle similar challenges in mice and non-human primates, and to begin to set the stage to address these questions in humans.
Finally, for all the advances to happen efficiently and equitably, patient volunteers and their families must be invited to the table. Scientists and physician-scientists can ensure respectful and affirming community engagement by partnering with patient advocacy groups, meeting with community partners, and offering accessibility through outreach and educational events130,131,132. These practices require preparation of team leaders and members and the appropriate skills to perform these tasks. Engagement involves creating trust, allowing dynamic communication, and ultimately giving back to the community, through the discovery and increased awareness of the neuroscience and neuropathology133. Healing happens with the potential for therapy but also with the acknowledgement and sharing of gained knowledge.
We propose that the era of human neuroscience has begun and will lead to transformative discoveries in fundamental and clinical neuroscience over the next decade. Critical to its success will be the ability to build a human neuroscience community that embraces the principles of data sharing that have enabled the genomics community to succeed.
References
-
Burns, S. P., Xing, D. & Shapley, R. M. Comparisons of the dynamics of local field potential and multiunit activity signals in macaque visual cortex. J. Neurosci. 30, 13739–13749 (2010).
-
Ahmadi, N., Constandinou, T. G. & Bouganis, C. S. Inferring entire spiking activity from local field potentials. Sci. Rep. 11, 19045 (2021).
-
Tseng, H. A. & Han, X. Distinct spiking patterns of excitatory and inhibitory neurons and LFP oscillations in prefrontal cortex during sensory discrimination. Front. Physiol. 12, 618307 (2021).
-
Cho, K. K. et al. Gamma rhythms link prefrontal interneuron dysfunction with cognitive inflexibility in Dlx5/6+/− mice. Neuron 85, 1332–1343 (2015).
-
Helfrich, R. F. & Knight, R. T. Oscillatory dynamics of prefrontal cognitive control. Trends Cogn. Sci. 20, 916–930 (2016).
-
Kamigaki, T. & Dan, Y. Delay activity of specific prefrontal interneuron subtypes modulates memory-guided behavior. Nat. Neurosci. 20, 854–863 (2017).
-
Pinto, L. & Dan, Y. Cell-type-specific activity in prefrontal cortex during goal-directed behavior. Neuron 87, 437–450 (2015).
-
Lagler, M. et al. Divisions of identified parvalbumin-expressing basket cells during working memory-guided decision making. Neuron 91, 1390–1401 (2016).
-
Kvitsiani, D. et al. Distinct behavioural and network correlates of two interneuron types in prefrontal cortex. Nature 498, 363–366 (2013).
-
Liebe, S., Hoerzer, G. M., Logothetis, N. K. & Rainer, G. Theta coupling between V4 and prefrontal cortex predicts visual short-term memory performance. Nat. Neurosci. 15, 456–462 (2012).
[Article](https://doi.org/10.1038%2Fnn.3038) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC38XhtlOrs7o%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=22286175) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Theta%20coupling%20between%20V4%20and%20prefrontal%20cortex%20predicts%20visual%20short-term%20memory%20performance&journal=Nat.%20Neurosci.&doi=10.1038%2Fnn.3038&volume=15&pages=456-462&publication_year=2012&author=Liebe%2CS&author=Hoerzer%2CGM&author=Logothetis%2CNK&author=Rainer%2CG)
- Buschman, T. J., Denovellis, E. L., Diogo, C., Bullock, D. & Miller, E. K. Synchronous oscillatory neural ensembles for rules in the prefrontal cortex. Neuron 76, 838–846 (2012).
[Article](https://doi.org/10.1016%2Fj.neuron.2012.09.029) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC38Xhsleju7fP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=23177967) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3907768) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Synchronous%20oscillatory%20neural%20ensembles%20for%20rules%20in%20the%20prefrontal%20cortex&journal=Neuron&doi=10.1016%2Fj.neuron.2012.09.029&volume=76&pages=838-846&publication_year=2012&author=Buschman%2CTJ&author=Denovellis%2CEL&author=Diogo%2CC&author=Bullock%2CD&author=Miller%2CEK)
- Hsieh, H. L., Wong, Y. T., Pesaran, B. & Shanechi, M. M. Multiscale modeling and decoding algorithms for spike-field activity. J. Neural Eng. 16, 016018 (2019).
[Article](https://doi.org/10.1088%2F1741-2552%2Faaeb1a) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2019JNEng..16a6018H) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30523833) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Multiscale%20modeling%20and%20decoding%20algorithms%20for%20spike-field%20activity&journal=J.%20Neural%20Eng.&doi=10.1088%2F1741-2552%2Faaeb1a&volume=16&publication_year=2019&author=Hsieh%2CHL&author=Wong%2CYT&author=Pesaran%2CB&author=Shanechi%2CMM)
- Fu, Z. et al. The geometry of domain-general performance monitoring in the human medial frontal cortex. Science 376, eabm9922 (2022). This study is an example of using Behnke–Fried electrodes to collect single neuron activity in large patient cohorts.
[Article](https://doi.org/10.1126%2Fscience.abm9922) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhsVSlur%2FP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35511978) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9282918) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20geometry%20of%20domain-general%20performance%20monitoring%20in%20the%20human%20medial%20frontal%20cortex&journal=Science&doi=10.1126%2Fscience.abm9922&volume=376&publication_year=2022&author=Fu%2CZ)
- Steinmetz, N. A., Koch, C., Harris, K. D. & Carandini, M. Challenges and opportunities for large-scale electrophysiology with Neuropixels probes. Curr. Opin. Neurobiol. 50, 92–100 (2018).
[Article](https://doi.org/10.1016%2Fj.conb.2018.01.009) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1cXhvFGjsrk%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29444488) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5999351) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Challenges%20and%20opportunities%20for%20large-scale%20electrophysiology%20with%20Neuropixels%20probes&journal=Curr.%20Opin.%20Neurobiol.&doi=10.1016%2Fj.conb.2018.01.009&volume=50&pages=92-100&publication_year=2018&author=Steinmetz%2CNA&author=Koch%2CC&author=Harris%2CKD&author=Carandini%2CM)
- Chung, J. E. et al. High-density single-unit human cortical recordings using the Neuropixels probe. Neuron 110, 2409–2421.e3 (2022).
[Article](https://doi.org/10.1016%2Fj.neuron.2022.05.007) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhsFWgsLzK) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35679860) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=High-density%20single-unit%20human%20cortical%20recordings%20using%20the%20Neuropixels%20probe&journal=Neuron&doi=10.1016%2Fj.neuron.2022.05.007&volume=110&pages=2409-2421.e3&publication_year=2022&author=Chung%2CJE)
- Khanna, A. R. et al. Single-neuronal elements of speech production in humans. Nature 626, 603–610 (2024).
[Article](https://doi.org/10.1038%2Fs41586-023-06982-w) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB2cXisFWqs7w%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=38297120) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10866697) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-neuronal%20elements%20of%20speech%20production%20in%20humans&journal=Nature&doi=10.1038%2Fs41586-023-06982-w&volume=626&pages=603-610&publication_year=2024&author=Khanna%2CAR)
- Leonard, M. K. et al. This study highlights use of Neuropixels for investigating columnar processing in human speech encoding. Nature 626, 593–602 (2024). This study highlights the use of Neuropixels probes for investigating columnar processing in human speech encoding.
[Article](https://doi.org/10.1038%2Fs41586-023-06839-2) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXis1Sks7vE) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=38093008) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=This%20study%20highlights%20use%20of%20Neuropixels%20for%20investigating%20columnar%20processing%20in%20human%20speech%20encoding&journal=Nature&doi=10.1038%2Fs41586-023-06839-2&volume=626&pages=593-602&publication_year=2024&author=Leonard%2CMK)
-
Trautmann, E. M. et al. Large-scale high-density brain-wide neural recording in nonhuman primates. Preprint at bioRxiv https://doi.org/10.1101/2023.02.01.526664 (2023).
-
Tischbirek, C. H. et al. In vivo functional mapping of a cortical column at single-neuron resolution. Cell Rep. 27, 1319–1326.e5 (2019).
[Article](https://doi.org/10.1016%2Fj.celrep.2019.04.007) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXovVCnsr8%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31042460) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=In%20vivo%20functional%20mapping%20of%20a%20cortical%20column%20at%20single-neuron%20resolution&journal=Cell%20Rep.&doi=10.1016%2Fj.celrep.2019.04.007&volume=27&pages=1319-1326.e5&publication_year=2019&author=Tischbirek%2CCH)
- Khodagholy, D. et al. NeuroGrid: recording action potentials from the surface of the brain. Nat. Neurosci. 18, 310–315 (2015).
[Article](https://doi.org/10.1038%2Fnn.3905) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2cXitFKltLjI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=25531570) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=NeuroGrid%3A%20recording%20action%20potentials%20from%20the%20surface%20of%20the%20brain&journal=Nat.%20Neurosci.&doi=10.1038%2Fnn.3905&volume=18&pages=310-315&publication_year=2015&author=Khodagholy%2CD)
- Tchoe, Y. et al. Human brain mapping with multithousand-channel PtNRGrids resolves spatiotemporal dynamics. Sci. Transl. Med. 14, eabj1441 (2022).
[Article](https://doi.org/10.1126%2Fscitranslmed.abj1441) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35044788) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9650779) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Human%20brain%20mapping%20with%20multithousand-channel%20PtNRGrids%20resolves%20spatiotemporal%20dynamics&journal=Sci.%20Transl.%20Med.&doi=10.1126%2Fscitranslmed.abj1441&volume=14&publication_year=2022&author=Tchoe%2CY)
- Paulk, A. C. et al. Microscale physiological events on the human cortical surface. Cereb. Cortex 31, 3678–3700 (2021).
[Article](https://doi.org/10.1093%2Fcercor%2Fbhab040) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33749727) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8258438) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Microscale%20physiological%20events%20on%20the%20human%20cortical%20surface&journal=Cereb.%20Cortex&doi=10.1093%2Fcercor%2Fbhab040&volume=31&pages=3678-3700&publication_year=2021&author=Paulk%2CAC)
- Paulk, A. C. et al. Large-scale neural recordings with single neuron resolution using Neuropixels probes in human cortex. Nat. Neurosci. 25, 252–263 (2022). Refs. 15 and 23 were among the first studies to demonstrate the use of Neuropixels probes in human cortex.
[Article](https://doi.org/10.1038%2Fs41593-021-00997-0) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XisFKrt7c%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35102333) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Large-scale%20neural%20recordings%20with%20single%20neuron%20resolution%20using%20Neuropixels%20probes%20in%20human%20cortex&journal=Nat.%20Neurosci.&doi=10.1038%2Fs41593-021-00997-0&volume=25&pages=252-263&publication_year=2022&author=Paulk%2CAC)
- Bugeon, S. et al. A transcriptomic axis predicts state modulation of cortical interneurons. Nature 607, 330–338 (2022).
[Article](https://doi.org/10.1038%2Fs41586-022-04915-7) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2022Natur.607..330B) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xhslent7%2FI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35794483) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9279161) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20transcriptomic%20axis%20predicts%20state%20modulation%20of%20cortical%20interneurons&journal=Nature&doi=10.1038%2Fs41586-022-04915-7&volume=607&pages=330-338&publication_year=2022&author=Bugeon%2CS)
- Pollen, A. A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
[Article](https://doi.org/10.1038%2Fnbt.2967) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2cXht1Gqt7zJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=25086649) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4191988) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Low-coverage%20single-cell%20mRNA%20sequencing%20reveals%20cellular%20heterogeneity%20and%20activated%20signaling%20pathways%20in%20developing%20cerebral%20cortex&journal=Nat.%20Biotechnol.&doi=10.1038%2Fnbt.2967&volume=32&pages=1053-1058&publication_year=2014&author=Pollen%2CAA)
- Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017). This study applies next-generation single-cell RNA sequencing technologies to profile human cortical cell atlases in brain development.
[Article](https://doi.org/10.1126%2Fscience.aap8809) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2017Sci...358.1318N) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2sXhvFWht7vO) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29217575) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5991609) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Spatiotemporal%20gene%20expression%20trajectories%20reveal%20developmental%20hierarchies%20of%20the%20human%20cortex&journal=Science&doi=10.1126%2Fscience.aap8809&volume=358&pages=1318-1323&publication_year=2017&author=Nowakowski%2CTJ)
- Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
[Article](https://doi.org/10.1073%2Fpnas.1507125112) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2015PNAS..112.7285D) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2MXosVKrurg%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=26060301) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4466750) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20survey%20of%20human%20brain%20transcriptome%20diversity%20at%20the%20single%20cell%20level&journal=Proc.%20Natl%20Acad.%20Sci.%20USA&doi=10.1073%2Fpnas.1507125112&volume=112&pages=7285-7290&publication_year=2015&author=Darmanis%2CS)
- Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015). This study introduces a key method in development of next-generation single-cell RNA sequencing technologies.
[Article](https://doi.org/10.1016%2Fj.cell.2015.05.002) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2MXpt1Sgt7o%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=26000488) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4481139) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Highly%20parallel%20genome-wide%20expression%20profiling%20of%20individual%20cells%20using%20nanoliter%20droplets&journal=Cell&doi=10.1016%2Fj.cell.2015.05.002&volume=161&pages=1202-1214&publication_year=2015&author=Macosko%2CEZ)
-
Langlieb, J. et al. The molecular cytoarchitecture of the adult mouse brain. Nature 624, 333–342 (2023).
-
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023). This study uses single-cell RNA sequencing and spatial transcriptomics to map the entire mouse brain.
[Article](https://doi.org/10.1038%2Fs41586-023-06812-z) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2023Natur.624..317Y) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXis1SksLfJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=38092916) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10719114) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20high-resolution%20transcriptomic%20and%20spatial%20atlas%20of%20cell%20types%20in%20the%20whole%20mouse%20brain&journal=Nature&doi=10.1038%2Fs41586-023-06812-z&volume=624&pages=317-332&publication_year=2023&author=Yao%2CZ)
- Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023). This study provides an initial draft of a cell atlas of the human brain.
[Article](https://doi.org/10.1126%2Fscience.add7046) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXitFygtbbF) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37824663) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Transcriptomic%20diversity%20of%20cell%20types%20across%20the%20adult%20human%20brain&journal=Science&doi=10.1126%2Fscience.add7046&volume=382&publication_year=2023&author=Siletti%2CK)
- Clark, I. C. et al. Microfluidics-free single-cell genomics with templated emulsification. Nat. Biotechnol. 41, 1557–1566 (2023).
[Article](https://doi.org/10.1038%2Fs41587-023-01685-z) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXkslOru7o%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36879006) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10635830) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Microfluidics-free%20single-cell%20genomics%20with%20templated%20emulsification&journal=Nat.%20Biotechnol.&doi=10.1038%2Fs41587-023-01685-z&volume=41&pages=1557-1566&publication_year=2023&author=Clark%2CIC)
- Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 111–119 (2021). This study is an example that highlights unique cell-type-specific transcriptomic and physiologic properties of human neurons compared to model systems.
[Article](https://doi.org/10.1038%2Fs41586-021-03465-8) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021Natur.598..111B) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXit1WqsbjM) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34616062) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8494640) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Comparative%20cellular%20analysis%20of%20motor%20cortex%20in%20human%2C%20marmoset%20and%20mouse&journal=Nature&doi=10.1038%2Fs41586-021-03465-8&volume=598&pages=111-119&publication_year=2021&author=Bakken%2CTE)
- Luo, C. et al. Single-cell methylomes identify neuronal subtypes and regulatory elements in mammalian cortex. Science 357, 600–604 (2017).
[Article](https://doi.org/10.1126%2Fscience.aan3351) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2017Sci...357..600L) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2sXhtlehur%2FE) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=28798132) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5570439) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-cell%20methylomes%20identify%20neuronal%20subtypes%20and%20regulatory%20elements%20in%20mammalian%20cortex&journal=Science&doi=10.1126%2Fscience.aan3351&volume=357&pages=600-604&publication_year=2017&author=Luo%2CC)
- Ziffra, R. S. et al. Single-cell epigenomics reveals mechanisms of human cortical development. Nature 598, 205–213 (2021).
[Article](https://doi.org/10.1038%2Fs41586-021-03209-8) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021Natur.598..205Z) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXit1WqsbvE) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34616060) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8494642) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-cell%20epigenomics%20reveals%20mechanisms%20of%20human%20cortical%20development&journal=Nature&doi=10.1038%2Fs41586-021-03209-8&volume=598&pages=205-213&publication_year=2021&author=Ziffra%2CRS)
- Luo, C. et al. Single nucleus multi-omics identifies human cortical cell regulatory genome diversity. Cell Genomics 2, 100107 (2022).
[Article](https://doi.org/10.1016%2Fj.xgen.2022.100107) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhvFCktrfJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35419551) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9004682) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single%20nucleus%20multi-omics%20identifies%20human%20cortical%20cell%20regulatory%20genome%20diversity&journal=Cell%20Genomics&doi=10.1016%2Fj.xgen.2022.100107&volume=2&publication_year=2022&author=Luo%2CC)
- Gatto, L. et al. Initial recommendations for performing, benchmarking and reporting single-cell proteomics experiments. Nat. Methods 20, 375–386 (2023).
[Article](https://doi.org/10.1038%2Fs41592-023-01785-3) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXkt1ymtrk%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36864200) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10130941) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Initial%20recommendations%20for%20performing%2C%20benchmarking%20and%20reporting%20single-cell%20proteomics%20experiments&journal=Nat.%20Methods&doi=10.1038%2Fs41592-023-01785-3&volume=20&pages=375-386&publication_year=2023&author=Gatto%2CL)
- Alon, S. et al. Expansion sequencing: spatially precise in situ transcriptomics in intact biological systems. Science 371, eaax2656 (2021).
[Article](https://doi.org/10.1126%2Fscience.aax2656) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXivVSisL0%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33509999) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7900882) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Expansion%20sequencing%3A%20spatially%20precise%20in%20situ%20transcriptomics%20in%20intact%20biological%20systems&journal=Science&doi=10.1126%2Fscience.aax2656&volume=371&publication_year=2021&author=Alon%2CS)
- Niu, M. et al. Droplet-based transcriptome profiling of individual synapses. Nat. Biotechnol. 41, 1332–1344 (2023).
[Article](https://doi.org/10.1038%2Fs41587-022-01635-1) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXhslKitbc%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36646931) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Droplet-based%20transcriptome%20profiling%20of%20individual%20synapses&journal=Nat.%20Biotechnol.&doi=10.1038%2Fs41587-022-01635-1&volume=41&pages=1332-1344&publication_year=2023&author=Niu%2CM)
- Zeng, H. What is a cell type and how to define it. Cell 185, 2739–2755 (2022).
[Article](https://doi.org/10.1016%2Fj.cell.2022.06.031) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhvV2lu77P) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35868277) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9342916) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=What%20is%20a%20cell%20type%20and%20how%20to%20define%20it&journal=Cell&doi=10.1016%2Fj.cell.2022.06.031&volume=185&pages=2739-2755&publication_year=2022&author=Zeng%2CH)
- Fang, R. et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56–62 (2022).
[Article](https://doi.org/10.1126%2Fscience.abm1741) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2022Sci...377...56F) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xhslars7%2FM) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35771910) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9262715) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Conservation%20and%20divergence%20of%20cortical%20cell%20organization%20in%20human%20and%20mouse%20revealed%20by%20MERFISH&journal=Science&doi=10.1126%2Fscience.abm1741&volume=377&pages=56-62&publication_year=2022&author=Fang%2CR)
- Zhang, D. et al. Spatial epigenome-transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
[Article](https://doi.org/10.1038%2Fs41586-023-05795-1) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2023Natur.616..113Z) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXltlKiurk%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36922587) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10076218) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Spatial%20epigenome-transcriptome%20co-profiling%20of%20mammalian%20tissues&journal=Nature&doi=10.1038%2Fs41586-023-05795-1&volume=616&pages=113-122&publication_year=2023&author=Zhang%2CD)
- Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
[Article](https://doi.org/10.1126%2Fscience.aat5691) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29930089) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6339868) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Three-dimensional%20intact-tissue%20sequencing%20of%20single-cell%20transcriptional%20states&journal=Science&doi=10.1126%2Fscience.aat5691&volume=361&publication_year=2018&author=Wang%2CX)
- Glasser, M. F. et al. A multi-modal parcellation of human cerebral cortex. Nature 536, 171–178 (2016).
[Article](https://doi.org/10.1038%2Fnature18933) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2016Natur.536..171G) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC28Xht1GrurnK) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=27437579) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4990127) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20multi-modal%20parcellation%20of%20human%20cerebral%20cortex&journal=Nature&doi=10.1038%2Fnature18933&volume=536&pages=171-178&publication_year=2016&author=Glasser%2CMF)
-
Liharska, L. E. et al. A study of gene expression in the living human brain. Preprint at medRxiv https://doi.org/10.1101/2023.04.21.23288916 (2023).
-
Schneider, A. et al. Transcriptomic cell type structures in vivo neuronal activity across multiple timescales. Cell Rep. 42, 112318 (2023).
[Article](https://doi.org/10.1016%2Fj.celrep.2023.112318) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXmt1SmsLw%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36995938) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10539488) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Transcriptomic%20cell%20type%20structures%20in%20vivo%20neuronal%20activity%20across%20multiple%20timescales&journal=Cell%20Rep.&doi=10.1016%2Fj.celrep.2023.112318&volume=42&publication_year=2023&author=Schneider%2CA)
- Ting, J. T. et al. A robust ex vivo experimental platform for molecular-genetic dissection of adult human neocortical cell types and circuits. Sci. Rep. 8, 8407 (2018).
[Article](https://doi.org/10.1038%2Fs41598-018-26803-9) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2018NatSR...8.8407T) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29849137) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5976666) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20robust%20ex%20vivo%20experimental%20platform%20for%20molecular-genetic%20dissection%20of%20adult%20human%20neocortical%20cell%20types%20and%20circuits&journal=Sci.%20Rep.&doi=10.1038%2Fs41598-018-26803-9&volume=8&publication_year=2018&author=Ting%2CJT)
- Beaulieu-Laroche, L. et al. Enhanced dendritic compartmentalization in human cortical neurons. Cell 175, 643–651.e14 (2018).
[Article](https://doi.org/10.1016%2Fj.cell.2018.08.045) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1cXhvFKisr3J) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30340039) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6197488) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Enhanced%20dendritic%20compartmentalization%20in%20human%20cortical%20neurons&journal=Cell&doi=10.1016%2Fj.cell.2018.08.045&volume=175&pages=643-651.e14&publication_year=2018&author=Beaulieu-Laroche%2CL)
- Moradi Chameh, H. et al. Diversity amongst human cortical pyramidal neurons revealed via their sag currents and frequency preferences. Nat. Commun. 12, 2497 (2021).
[Article](https://doi.org/10.1038%2Fs41467-021-22741-9) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021NatCo..12.2497M) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXhtVGlsLbI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33941783) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8093195) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Diversity%20amongst%20human%20cortical%20pyramidal%20neurons%20revealed%20via%20their%20sag%20currents%20and%20frequency%20preferences&journal=Nat.%20Commun.&doi=10.1038%2Fs41467-021-22741-9&volume=12&publication_year=2021&author=Moradi%20Chameh%2CH)
- Kalmbach, B. E. et al. h-Channels contribute to divergent intrinsic membrane properties of supragranular pyramidal neurons in human versus mouse cerebral cortex. Neuron 100, 1194–1208.e5 (2018).
[Article](https://doi.org/10.1016%2Fj.neuron.2018.10.012) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1cXitVyhsbbL) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30392798) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6447369) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=h-Channels%20contribute%20to%20divergent%20intrinsic%20membrane%20properties%20of%20supragranular%20pyramidal%20neurons%20in%20human%20versus%20mouse%20cerebral%20cortex&journal=Neuron&doi=10.1016%2Fj.neuron.2018.10.012&volume=100&pages=1194-1208.e5&publication_year=2018&author=Kalmbach%2CBE)
- Beaulieu-Laroche, L. et al. Allometric rules for mammalian cortical layer 5 neuron biophysics. Nature 600, 274–278 (2021).
[Article](https://doi.org/10.1038%2Fs41586-021-04072-3) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021Natur.600..274B) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXisVOltLbI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34759318) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Allometric%20rules%20for%20mammalian%20cortical%20layer%205%20neuron%20biophysics&journal=Nature&doi=10.1038%2Fs41586-021-04072-3&volume=600&pages=274-278&publication_year=2021&author=Beaulieu-Laroche%2CL)
- Berto, S. et al. Gene-expression correlates of the oscillatory signatures supporting human episodic memory encoding. Nat. Neurosci. 24, 554–564 (2021).
[Article](https://doi.org/10.1038%2Fs41593-021-00803-x) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXmtVSrtrs%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33686299) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8016736) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Gene-expression%20correlates%20of%20the%20oscillatory%20signatures%20supporting%20human%20episodic%20memory%20encoding&journal=Nat.%20Neurosci.&doi=10.1038%2Fs41593-021-00803-x&volume=24&pages=554-564&publication_year=2021&author=Berto%2CS)
- Mahalingam, G. et al. A scalable and modular automated pipeline for stitching of large electron microscopy datasets. eLife 11, e76534 (2022).
[Article](https://doi.org/10.7554%2FeLife.76534) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXltVCmsb8%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35880860) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9427110) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20scalable%20and%20modular%20automated%20pipeline%20for%20stitching%20of%20large%20electron%20microscopy%20datasets&journal=eLife&doi=10.7554%2FeLife.76534&volume=11&publication_year=2022&author=Mahalingam%2CG)
- Chen, X. et al. High-throughput mapping of long-range neuronal projection using in situ sequencing. Cell 179, 772–786.e19 (2019).
[Article](https://doi.org/10.1016%2Fj.cell.2019.09.023) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXitVSls7bJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31626774) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7836778) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=High-throughput%20mapping%20of%20long-range%20neuronal%20projection%20using%20in%20situ%20sequencing&journal=Cell&doi=10.1016%2Fj.cell.2019.09.023&volume=179&pages=772-786.e19&publication_year=2019&author=Chen%2CX)
- Foss-Feig, J. H. et al. Searching for cross-diagnostic convergence: neural mechanisms governing excitation and inhibition balance in schizophrenia and autism spectrum disorders. Biol. Psychiatry 81, 848–861 (2017).
[Article](https://doi.org/10.1016%2Fj.biopsych.2017.03.005) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=28434615) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5436134) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Searching%20for%20cross-diagnostic%20convergence%3A%20neural%20mechanisms%20governing%20excitation%20and%20inhibition%20balance%20in%20schizophrenia%20and%20autism%20spectrum%20disorders&journal=Biol.%20Psychiatry&doi=10.1016%2Fj.biopsych.2017.03.005&volume=81&pages=848-861&publication_year=2017&author=Foss-Feig%2CJH)
- Krystal, J. H. et al. Impaired tuning of neural ensembles and the pathophysiology of schizophrenia: a translational and computational neuroscience perspective. Biol. Psychiatry 81, 874–885 (2017).
[Article](https://doi.org/10.1016%2Fj.biopsych.2017.01.004) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2sXms12lsLs%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=28434616) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5407407) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Impaired%20tuning%20of%20neural%20ensembles%20and%20the%20pathophysiology%20of%20schizophrenia%3A%20a%20translational%20and%20computational%20neuroscience%20perspective&journal=Biol.%20Psychiatry&doi=10.1016%2Fj.biopsych.2017.01.004&volume=81&pages=874-885&publication_year=2017&author=Krystal%2CJH)
- Opris, I. & Casanova, M. F. Prefrontal cortical minicolumn: from executive control to disrupted cognitive processing. Brain 137, 1863–1875 (2014).
[Article](https://doi.org/10.1093%2Fbrain%2Fawt359) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=24531625) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4065017) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Prefrontal%20cortical%20minicolumn%3A%20from%20executive%20control%20to%20disrupted%20cognitive%20processing&journal=Brain&doi=10.1093%2Fbrain%2Fawt359&volume=137&pages=1863-1875&publication_year=2014&author=Opris%2CI&author=Casanova%2CMF)
- Li, Q. et al. Multimodal charting of molecular and functional cell states via in situ electro-sequencing. Cell 186, 2002–2017.e21 (2023).
[Article](https://doi.org/10.1016%2Fj.cell.2023.03.023) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXotFCkt7Y%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37080201) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Multimodal%20charting%20of%20molecular%20and%20functional%20cell%20states%20via%20in%20situ%20electro-sequencing&journal=Cell&doi=10.1016%2Fj.cell.2023.03.023&volume=186&pages=2002-2017.e21&publication_year=2023&author=Li%2CQ)
- Billeh, Y. N. et al. Systematic integration of structural and functional data into multi-scale models of mouse primary visual cortex. Neuron 106, 388–403.e18 (2020).
[Article](https://doi.org/10.1016%2Fj.neuron.2020.01.040) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXksFaqu74%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32142648) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Systematic%20integration%20of%20structural%20and%20functional%20data%20into%20multi-scale%20models%20of%20mouse%20primary%20visual%20cortex&journal=Neuron&doi=10.1016%2Fj.neuron.2020.01.040&volume=106&pages=388-403.e18&publication_year=2020&author=Billeh%2CYN)
- Buchin, A. et al. Multi-modal characterization and simulation of human epileptic circuitry. Cell Rep. 41, 111873 (2022).
[Article](https://doi.org/10.1016%2Fj.celrep.2022.111873) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xjt1SnsL7J) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36577383) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9841067) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Multi-modal%20characterization%20and%20simulation%20of%20human%20epileptic%20circuitry&journal=Cell%20Rep.&doi=10.1016%2Fj.celrep.2022.111873&volume=41&publication_year=2022&author=Buchin%2CA)
- Winkler, E. A. et al. A single-cell atlas of the normal and malformed human brain vasculature. Science 375, eabi7377 (2022). This study describes an example of how an ex vivo platform can be utilized to study non-neuronal cell types in human brain and disease.
[Article](https://doi.org/10.1126%2Fscience.abi7377) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xmtl2gtLc%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35084939) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8995178) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20single-cell%20atlas%20of%20the%20normal%20and%20malformed%20human%20brain%20vasculature&journal=Science&doi=10.1126%2Fscience.abi7377&volume=375&publication_year=2022&author=Winkler%2CEA)
- Eugène, E. et al. An organotypic brain slice preparation from adult patients with temporal lobe epilepsy. J. Neurosci. Methods 235, 234–244 (2014).
[Article](https://doi.org/10.1016%2Fj.jneumeth.2014.07.009) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=25064188) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4426207) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=An%20organotypic%20brain%20slice%20preparation%20from%20adult%20patients%20with%20temporal%20lobe%20epilepsy&journal=J.%20Neurosci.%20Methods&doi=10.1016%2Fj.jneumeth.2014.07.009&volume=235&pages=234-244&publication_year=2014&author=Eug%C3%A8ne%2CE)
- Schwarz, N. et al. Long-term adult human brain slice cultures as a model system to study human CNS circuitry and disease. eLife 8, e48417 (2019).
[Article](https://doi.org/10.7554%2FeLife.48417) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXhtlaltL%2FP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31498083) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6733599) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Long-term%20adult%20human%20brain%20slice%20cultures%20as%20a%20model%20system%20to%20study%20human%20CNS%20circuitry%20and%20disease&journal=eLife&doi=10.7554%2FeLife.48417&volume=8&publication_year=2019&author=Schwarz%2CN)
- Mich, J. K. et al. Functional enhancer elements drive subclass-selective expression from mouse to primate neocortex. Cell Rep. 34, 108754 (2021).
[Article](https://doi.org/10.1016%2Fj.celrep.2021.108754) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXnvFCiu7o%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33789096) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8163032) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Functional%20enhancer%20elements%20drive%20subclass-selective%20expression%20from%20mouse%20to%20primate%20neocortex&journal=Cell%20Rep.&doi=10.1016%2Fj.celrep.2021.108754&volume=34&publication_year=2021&author=Mich%2CJK)
- Zhu, D. et al. Optimal trade-off control in machine learning-based library design, with application to adeno-associated virus (AAV) for gene therapy. Sci. Adv. 10, eadj3786 (2024).
[Article](https://doi.org/10.1126%2Fsciadv.adj3786) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=38266077) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10807795) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Optimal%20trade-off%20control%20in%20machine%20learning-based%20library%20design%2C%20with%20application%20to%20adeno-associated%20virus%20%28AAV%29%20for%20gene%20therapy&journal=Sci.%20Adv.&doi=10.1126%2Fsciadv.adj3786&volume=10&publication_year=2024&author=Zhu%2CD)
- Mercuri, E., Pera, M. C., Scoto, M., Finkel, R. & Muntoni, F. Spinal muscular atrophy—insights and challenges in the treatment era. Nat. Rev. Neurol. 16, 706–715 (2020).
[Article](https://doi.org/10.1038%2Fs41582-020-00413-4) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXitlWku7rM) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33057172) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Spinal%20muscular%20atrophy%E2%80%94insights%20and%20challenges%20in%20the%20treatment%20era&journal=Nat.%20Rev.%20Neurol.&doi=10.1038%2Fs41582-020-00413-4&volume=16&pages=706-715&publication_year=2020&author=Mercuri%2CE&author=Pera%2CMC&author=Scoto%2CM&author=Finkel%2CR&author=Muntoni%2CF)
- Neuner, S. M., Tcw, J. & Goate, A. M. Genetic architecture of Alzheimer’s disease. Neurobiol. Dis. 143, 104976 (2020).
[Article](https://doi.org/10.1016%2Fj.nbd.2020.104976) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXhtlSktbfN) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32565066) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7409822) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Genetic%20architecture%20of%20Alzheimer%E2%80%99s%20disease&journal=Neurobiol.%20Dis.&doi=10.1016%2Fj.nbd.2020.104976&volume=143&publication_year=2020&author=Neuner%2CSM&author=Tcw%2CJ&author=Goate%2CAM)
- Tabrizi, S. J., Flower, M. D., Ross, C. A. & Wild, E. J. Huntington disease: new insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. 16, 529–546 (2020).
[Article](https://doi.org/10.1038%2Fs41582-020-0389-4) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32796930) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Huntington%20disease%3A%20new%20insights%20into%20molecular%20pathogenesis%20and%20therapeutic%20opportunities&journal=Nat.%20Rev.%20Neurol.&doi=10.1038%2Fs41582-020-0389-4&volume=16&pages=529-546&publication_year=2020&author=Tabrizi%2CSJ&author=Flower%2CMD&author=Ross%2CCA&author=Wild%2CEJ)
- Niestroj, L. M. et al. Epilepsy subtype-specific copy number burden observed in a genome-wide study of 17 458 subjects. Brain 143, 2106–2118 (2020).
[Article](https://doi.org/10.1093%2Fbrain%2Fawaa171) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32568404) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7364765) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Epilepsy%20subtype-specific%20copy%20number%20burden%20observed%20in%20a%20genome-wide%20study%20of%2017%E2%80%89458%20subjects&journal=Brain&doi=10.1093%2Fbrain%2Fawaa171&volume=143&pages=2106-2118&publication_year=2020&author=Niestroj%2CLM)
- Epi4K Consortiu. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
[Article](https://doi.org/10.1038%2Fnature12439) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2013Natur.501..217E) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=De%20novo%20mutations%20in%20epileptic%20encephalopathies&journal=Nature&doi=10.1038%2Fnature12439&volume=501&pages=217-221&publication_year=2013)
- Wang, T. et al. Integrated gene analyses of de novo variants from 46,612 trios with autism and developmental disorders. Proc. Natl Acad. Sci. USA 119, e2203491119 (2022).
[Article](https://doi.org/10.1073%2Fpnas.2203491119) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XjtVOhurnO) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36350923) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9674258) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Integrated%20gene%20analyses%20of%20de%20novo%20variants%20from%2046%2C612%20trios%20with%20autism%20and%20developmental%20disorders&journal=Proc.%20Natl%20Acad.%20Sci.%20USA&doi=10.1073%2Fpnas.2203491119&volume=119&publication_year=2022&author=Wang%2CT)
- Fu, J. M. et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat. Genet. 54, 1320–1331 (2022).
[Article](https://doi.org/10.1038%2Fs41588-022-01104-0) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XitFOksLzI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35982160) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9653013) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Rare%20coding%20variation%20provides%20insight%20into%20the%20genetic%20architecture%20and%20phenotypic%20context%20of%20autism&journal=Nat.%20Genet.&doi=10.1038%2Fs41588-022-01104-0&volume=54&pages=1320-1331&publication_year=2022&author=Fu%2CJM)
- Singh, T. et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature 604, 509–516 (2022).
[Article](https://doi.org/10.1038%2Fs41586-022-04556-w) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2022Natur.604..509S) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XpvVWjtrs%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35396579) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9805802) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Rare%20coding%20variants%20in%20ten%20genes%20confer%20substantial%20risk%20for%20schizophrenia&journal=Nature&doi=10.1038%2Fs41586-022-04556-w&volume=604&pages=509-516&publication_year=2022&author=Singh%2CT)
- Liu, D. et al. Schizophrenia risk conferred by rare protein-truncating variants is conserved across diverse human populations. Nat. Genet. 55, 369–376 (2023).
[Article](https://doi.org/10.1038%2Fs41588-023-01305-1) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXltFGmtLg%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36914870) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10011128) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Schizophrenia%20risk%20conferred%20by%20rare%20protein-truncating%20variants%20is%20conserved%20across%20diverse%20human%20populations&journal=Nat.%20Genet.&doi=10.1038%2Fs41588-023-01305-1&volume=55&pages=369-376&publication_year=2023&author=Liu%2CD)
- Lin, X. et al. Integrating genetic structural variations and whole-genome sequencing into clinical neurology. Neurol. Genet. 8, e200005 (2022).
[Article](https://doi.org/10.1212%2FNXG.0000000000200005) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37435434) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10331588) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Integrating%20genetic%20structural%20variations%20and%20whole-genome%20sequencing%20into%20clinical%20neurology&journal=Neurol.%20Genet.&doi=10.1212%2FNXG.0000000000200005&volume=8&publication_year=2022&author=Lin%2CX)
- Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
[Article](https://doi.org/10.1038%2Fnature11011) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2012Natur.485..242N) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC38XmtVerurs%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=22495311) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3613847) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Patterns%20and%20rates%20of%20exonic%20de%20novo%20mutations%20in%20autism%20spectrum%20disorders&journal=Nature&doi=10.1038%2Fnature11011&volume=485&pages=242-245&publication_year=2012&author=Neale%2CBM)
- Mitra, I. et al. Patterns of de novo tandem repeat mutations and their role in autism. Nature 589, 246–250 (2021).
[Article](https://doi.org/10.1038%2Fs41586-020-03078-7) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021Natur.589..246M) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXht1Omu74%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33442040) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7810352) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Patterns%20of%20de%20novo%20tandem%20repeat%20mutations%20and%20their%20role%20in%20autism&journal=Nature&doi=10.1038%2Fs41586-020-03078-7&volume=589&pages=246-250&publication_year=2021&author=Mitra%2CI)
- Nishioka, M., Bundo, M., Iwamoto, K. & Kato, T. Somatic mutations in the human brain: implications for psychiatric research. Mol. Psychiatry 24, 839–856 (2019).
[Article](https://doi.org/10.1038%2Fs41380-018-0129-y) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30087451) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Somatic%20mutations%20in%20the%20human%20brain%3A%20implications%20for%20psychiatric%20research&journal=Mol.%20Psychiatry&doi=10.1038%2Fs41380-018-0129-y&volume=24&pages=839-856&publication_year=2019&author=Nishioka%2CM&author=Bundo%2CM&author=Iwamoto%2CK&author=Kato%2CT)
- Rehen, S. K. et al. Constitutional aneuploidy in the normal human brain. J. Neurosci. 25, 2176–2180 (2005).
[Article](https://doi.org/10.1523%2FJNEUROSCI.4560-04.2005) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BD2MXisVOmtro%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=15745943) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6726097) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Constitutional%20aneuploidy%20in%20the%20normal%20human%20brain&journal=J.%20Neurosci.&doi=10.1523%2FJNEUROSCI.4560-04.2005&volume=25&pages=2176-2180&publication_year=2005&author=Rehen%2CSK)
- Wang, Y. et al. Comprehensive identification of somatic nucleotide variants in human brain tissue. Genome Biol. 22, 92 (2021).
[Article](https://link.springer.com/doi/10.1186/s13059-021-02285-3) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXht1KrtrrF) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33781308) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8006362) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Comprehensive%20identification%20of%20somatic%20nucleotide%20variants%20in%20human%20brain%20tissue&journal=Genome%20Biol.&doi=10.1186%2Fs13059-021-02285-3&volume=22&publication_year=2021&author=Wang%2CY)
- Luquette, L. J. et al. Single-cell genome sequencing of human neurons identifies somatic point mutation and indel enrichment in regulatory elements. Nat. Genet. 54, 1564–1571 (2022).
[Article](https://doi.org/10.1038%2Fs41588-022-01180-2) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XisV2nsrvO) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36163278) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9833626) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-cell%20genome%20sequencing%20of%20human%20neurons%20identifies%20somatic%20point%20mutation%20and%20indel%20enrichment%20in%20regulatory%20elements&journal=Nat.%20Genet.&doi=10.1038%2Fs41588-022-01180-2&volume=54&pages=1564-1571&publication_year=2022&author=Luquette%2CLJ)
- Chung, C. et al. Comprehensive multi-omic profiling of somatic mutations in malformations of cortical development. Nat. Genet. 55, 209–220 (2023).
[Article](https://doi.org/10.1038%2Fs41588-022-01276-9) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXhtV2lsrg%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36635388) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9961399) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Comprehensive%20multi-omic%20profiling%20of%20somatic%20mutations%20in%20malformations%20of%20cortical%20development&journal=Nat.%20Genet.&doi=10.1038%2Fs41588-022-01276-9&volume=55&pages=209-220&publication_year=2023&author=Chung%2CC)
- Hua, Y. & Crino, P. B. Single cell lineage analysis in human focal cortical dysplasia. Cereb. Cortex 13, 693–699 (2003).
[Article](https://doi.org/10.1093%2Fcercor%2F13.6.693) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=12764046) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single%20cell%20lineage%20analysis%20in%20human%20focal%20cortical%20dysplasia&journal=Cereb.%20Cortex&doi=10.1093%2Fcercor%2F13.6.693&volume=13&pages=693-699&publication_year=2003&author=Hua%2CY&author=Crino%2CPB)
- Rivière, J. B. et al. De novo germline and postzygotic mutations in AKT3, PIK3R2 and PIK3CA cause a spectrum of related megalencephaly syndromes. Nat. Genet. 44, 934–940 (2012).
[Article](https://doi.org/10.1038%2Fng.2331) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=22729224) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3408813) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=De%20novo%20germline%20and%20postzygotic%20mutations%20in%20AKT3%2C%20PIK3R2%20and%20PIK3CA%20cause%20a%20spectrum%20of%20related%20megalencephaly%20syndromes&journal=Nat.%20Genet.&doi=10.1038%2Fng.2331&volume=44&pages=934-940&publication_year=2012&author=Rivi%C3%A8re%2CJB)
- Poduri, A. et al. Somatic activation of AKT3 causes hemispheric developmental brain malformations. Neuron 74, 41–48 (2012).
[Article](https://doi.org/10.1016%2Fj.neuron.2012.03.010) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC38Xls1ehuro%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=22500628) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3460551) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Somatic%20activation%20of%20AKT3%20causes%20hemispheric%20developmental%20brain%20malformations&journal=Neuron&doi=10.1016%2Fj.neuron.2012.03.010&volume=74&pages=41-48&publication_year=2012&author=Poduri%2CA)
- Ribierre, T. et al. Second-hit mosaic mutation in mTORC1 repressor DEPDC5 causes focal cortical dysplasia-associated epilepsy. J. Clin. Invest. 128, 2452–2458 (2018).
[Article](https://doi.org/10.1172%2FJCI99384) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29708508) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5983335) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Second-hit%20mosaic%20mutation%20in%20mTORC1%20repressor%20DEPDC5%20causes%20focal%20cortical%20dysplasia-associated%20epilepsy&journal=J.%20Clin.%20Invest.&doi=10.1172%2FJCI99384&volume=128&pages=2452-2458&publication_year=2018&author=Ribierre%2CT)
- Baldassari, S. et al. Dissecting the genetic basis of focal cortical dysplasia: a large cohort study. Acta Neuropathol. 138, 885–900 (2019).
[Article](https://link.springer.com/doi/10.1007/s00401-019-02061-5) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXhs1eksb%2FJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31444548) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6851393) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Dissecting%20the%20genetic%20basis%20of%20focal%20cortical%20dysplasia%3A%20a%20large%20cohort%20study&journal=Acta%20Neuropathol.&doi=10.1007%2Fs00401-019-02061-5&volume=138&pages=885-900&publication_year=2019&author=Baldassari%2CS)
- Wirrell, E. C. et al. Optimizing the diagnosis and management of dravet syndrome: recommendations from a North American consensus panel. Pediatr. Neurol. 68, 18–34.e3 (2017).
[Article](https://doi.org/10.1016%2Fj.pediatrneurol.2017.01.025) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=28284397) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Optimizing%20the%20diagnosis%20and%20management%20of%20dravet%20syndrome%3A%20recommendations%20from%20a%20North%20American%20consensus%20panel&journal=Pediatr.%20Neurol.&doi=10.1016%2Fj.pediatrneurol.2017.01.025&volume=68&pages=18-34.e3&publication_year=2017&author=Wirrell%2CEC)
- Gratuze, M., Leyns, C. E. G. & Holtzman, D. M. New insights into the role of TREM2 in Alzheimer’s disease. Mol. Neurodegener. 13, 66 (2018).
[Article](https://link.springer.com/doi/10.1186/s13024-018-0298-9) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXhtFehsLvE) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30572908) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6302500) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=New%20insights%20into%20the%20role%20of%20TREM2%20in%20Alzheimer%E2%80%99s%20disease&journal=Mol.%20Neurodegener.&doi=10.1186%2Fs13024-018-0298-9&volume=13&publication_year=2018&author=Gratuze%2CM&author=Leyns%2CCEG&author=Holtzman%2CDM)
- Ulland, T. K. & Colonna, M. TREM2—a key player in microglial biology and Alzheimer disease. Nat. Rev. Neurol. 14, 667–675 (2018).
[Article](https://doi.org/10.1038%2Fs41582-018-0072-1) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1cXhvVeru7fI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30266932) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=TREM2%E2%80%94a%20key%20player%20in%20microglial%20biology%20and%20Alzheimer%20disease&journal=Nat.%20Rev.%20Neurol.&doi=10.1038%2Fs41582-018-0072-1&volume=14&pages=667-675&publication_year=2018&author=Ulland%2CTK&author=Colonna%2CM)
- Braasch, D. A., Liu, Y. & Corey, D. R. Antisense inhibition of gene expression in cells by oligonucleotides incorporating locked nucleic acids: effect of mRNA target sequence and chimera design. Nucleic Acids Res. 30, 5160–5167 (2002).
[Article](https://doi.org/10.1093%2Fnar%2Fgkf651) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BD38XptlOiu74%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=12466540) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC137965) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Antisense%20inhibition%20of%20gene%20expression%20in%20cells%20by%20oligonucleotides%20incorporating%20locked%20nucleic%20acids%3A%20effect%20of%20mRNA%20target%20sequence%20and%20chimera%20design&journal=Nucleic%20Acids%20Res.&doi=10.1093%2Fnar%2Fgkf651&volume=30&pages=5160-5167&publication_year=2002&author=Braasch%2CDA&author=Liu%2CY&author=Corey%2CDR)
- Bladen, C. L. et al. The TREAT-NMD DMD Global Database: analysis of more than 7,000 Duchenne muscular dystrophy mutations. Hum. Mutat. 36, 395–402 (2015).
[Article](https://doi.org/10.1002%2Fhumu.22758) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2MXlsFaktbg%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=25604253) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4405042) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20TREAT-NMD%20DMD%20Global%20Database%3A%20analysis%20of%20more%20than%207%2C000%20Duchenne%20muscular%20dystrophy%20mutations&journal=Hum.%20Mutat.&doi=10.1002%2Fhumu.22758&volume=36&pages=395-402&publication_year=2015&author=Bladen%2CCL)
- Finkel, R. S. et al. Treatment of infantile-onset spinal muscular atrophy with nusinersen: a phase 2, open-label, dose-escalation study. Lancet 388, 3017–3026 (2016).
[Article](https://doi.org/10.1016%2FS0140-6736%2816%2931408-8) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC28XitVegurfO) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=27939059) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Treatment%20of%20infantile-onset%20spinal%20muscular%20atrophy%20with%20nusinersen%3A%20a%20phase%202%2C%20open-label%2C%20dose-escalation%20study&journal=Lancet&doi=10.1016%2FS0140-6736%2816%2931408-8&volume=388&pages=3017-3026&publication_year=2016&author=Finkel%2CRS)
- Hampel, H. et al. The foundation and architecture of precision medicine in neurology and psychiatry. Trends Neurosci. 46, 176–198 (2023).
[Article](https://doi.org/10.1016%2Fj.tins.2022.12.004) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XjtF2qsr%2FF) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36642626) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10720395) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20foundation%20and%20architecture%20of%20precision%20medicine%20in%20neurology%20and%20psychiatry&journal=Trends%20Neurosci.&doi=10.1016%2Fj.tins.2022.12.004&volume=46&pages=176-198&publication_year=2023&author=Hampel%2CH)
- Knowles, J. K. et al. Precision medicine for genetic epilepsy on the horizon: recent advances, present challenges, and suggestions for continued progress. Epilepsia 63, 2461–2475 (2022).
[Article](https://doi.org/10.1111%2Fepi.17332) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35716052) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9561034) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Precision%20medicine%20for%20genetic%20epilepsy%20on%20the%20horizon%3A%20recent%20advances%2C%20present%20challenges%2C%20and%20suggestions%20for%20continued%20progress&journal=Epilepsia&doi=10.1111%2Fepi.17332&volume=63&pages=2461-2475&publication_year=2022&author=Knowles%2CJK)
- Varma, V. R. et al. Brain and blood metabolite signatures of pathology and progression in Alzheimer disease: a targeted metabolomics study. PLoS Med. 15, e1002482 (2018).
[Article](https://doi.org/10.1371%2Fjournal.pmed.1002482) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29370177) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5784884) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Brain%20and%20blood%20metabolite%20signatures%20of%20pathology%20and%20progression%20in%20Alzheimer%20disease%3A%20a%20targeted%20metabolomics%20study&journal=PLoS%20Med.&doi=10.1371%2Fjournal.pmed.1002482&volume=15&publication_year=2018&author=Varma%2CVR)
- Noori, A., Mezlini, A. M., Hyman, B. T., Serrano-Pozo, A. & Das, S. Systematic review and meta-analysis of human transcriptomics reveals neuroinflammation, deficient energy metabolism, and proteostasis failure across neurodegeneration. Neurobiol. Dis. 149, 105225 (2021).
[Article](https://doi.org/10.1016%2Fj.nbd.2020.105225) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXisVWnt7k%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33347974) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Systematic%20review%20and%20meta-analysis%20of%20human%20transcriptomics%20reveals%20neuroinflammation%2C%20deficient%20energy%20metabolism%2C%20and%20proteostasis%20failure%20across%20neurodegeneration&journal=Neurobiol.%20Dis.&doi=10.1016%2Fj.nbd.2020.105225&volume=149&publication_year=2021&author=Noori%2CA&author=Mezlini%2CAM&author=Hyman%2CBT&author=Serrano-Pozo%2CA&author=Das%2CS)
- Quazi, S. Artificial intelligence and machine learning in precision and genomic medicine. Med. Oncol. 39, 120 (2022).
[Article](https://link.springer.com/doi/10.1007/s12032-022-01711-1) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35704152) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9198206) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Artificial%20intelligence%20and%20machine%20learning%20in%20precision%20and%20genomic%20medicine&journal=Med.%20Oncol.&doi=10.1007%2Fs12032-022-01711-1&volume=39&publication_year=2022&author=Quazi%2CS)
- Ingusci, S., Verlengia, G., Soukupova, M., Zucchini, S. & Simonato, M. Gene therapy tools for brain diseases. Front. Pharmacol. 10, 724 (2019).
[Article](https://doi.org/10.3389%2Ffphar.2019.00724) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXksVarsbY%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31312139) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6613496) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Gene%20therapy%20tools%20for%20brain%20diseases&journal=Front.%20Pharmacol.&doi=10.3389%2Ffphar.2019.00724&volume=10&publication_year=2019&author=Ingusci%2CS&author=Verlengia%2CG&author=Soukupova%2CM&author=Zucchini%2CS&author=Simonato%2CM)
- Zhu, D., Schieferecke, A. J., Lopez, P. A. & Schaffer, D. V. Adeno-associated virus vector for central nervous system gene therapy. Trends Mol. Med. 27, 524–537 (2021).
[Article](https://doi.org/10.1016%2Fj.molmed.2021.03.010) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXhtlGmsLrP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33895085) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Adeno-associated%20virus%20vector%20for%20central%20nervous%20system%20gene%20therapy&journal=Trends%20Mol.%20Med.&doi=10.1016%2Fj.molmed.2021.03.010&volume=27&pages=524-537&publication_year=2021&author=Zhu%2CD&author=Schieferecke%2CAJ&author=Lopez%2CPA&author=Schaffer%2CDV)
- Sugarman, E. A. et al. Pan-ethnic carrier screening and prenatal diagnosis for spinal muscular atrophy: clinical laboratory analysis of >72,400 specimens. Eur. J. Hum. Genet. 20, 27–32 (2012).
[Article](https://doi.org/10.1038%2Fejhg.2011.134) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=21811307) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Pan-ethnic%20carrier%20screening%20and%20prenatal%20diagnosis%20for%20spinal%20muscular%20atrophy%3A%20clinical%20laboratory%20analysis%20of%20%3E72%2C400%20specimens&journal=Eur.%20J.%20Hum.%20Genet.&doi=10.1038%2Fejhg.2011.134&volume=20&pages=27-32&publication_year=2012&author=Sugarman%2CEA)
- Mendell, J. R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
[Article](https://doi.org/10.1056%2FNEJMoa1706198) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2sXhvFChur7K) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=29091557) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-dose%20gene-replacement%20therapy%20for%20spinal%20muscular%20atrophy&journal=N.%20Engl.%20J.%20Med.&doi=10.1056%2FNEJMoa1706198&volume=377&pages=1713-1722&publication_year=2017&author=Mendell%2CJR)
- Mendell, J. R. et al. Current clinical applications of in vivo gene therapy with AAVs. Mol. Ther. 29, 464–488 (2021).
[Article](https://doi.org/10.1016%2Fj.ymthe.2020.12.007) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXhtVGju7k%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33309881) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Current%20clinical%20applications%20of%20in%20vivo%20gene%20therapy%20with%20AAVs&journal=Mol.%20Ther.&doi=10.1016%2Fj.ymthe.2020.12.007&volume=29&pages=464-488&publication_year=2021&author=Mendell%2CJR)
- Chien, Y. H. et al. Efficacy and safety of AAV2 gene therapy in children with aromatic l-amino acid decarboxylase deficiency: an open-label, phase 1/2 trial. Lancet Child Adolesc. Health 1, 265–273 (2017).
[Article](https://doi.org/10.1016%2FS2352-4642%2817%2930125-6) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30169182) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Efficacy%20and%20safety%20of%20AAV2%20gene%20therapy%20in%20children%20with%20aromatic%20l-amino%20acid%20decarboxylase%20deficiency%3A%20an%20open-label%2C%20phase%201%2F2%20trial&journal=Lancet%20Child%20Adolesc.%20Health&doi=10.1016%2FS2352-4642%2817%2930125-6&volume=1&pages=265-273&publication_year=2017&author=Chien%2CYH)
- Leone, P. et al. Long-term follow-up after gene therapy for canavan disease. Sci. Transl. Med. 4, 165ra163 (2012).
[Article](https://doi.org/10.1126%2Fscitranslmed.3003454) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=23253610) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3794457) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Long-term%20follow-up%20after%20gene%20therapy%20for%20canavan%20disease&journal=Sci.%20Transl.%20Med.&doi=10.1126%2Fscitranslmed.3003454&volume=4&publication_year=2012&author=Leone%2CP)
- Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
[Article](https://doi.org/10.1038%2Fnn.4593) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC2sXhtVOjsb3I) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=28671695) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5529245) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Engineered%20AAVs%20for%20efficient%20noninvasive%20gene%20delivery%20to%20the%20central%20and%20peripheral%20nervous%20systems&journal=Nat.%20Neurosci.&doi=10.1038%2Fnn.4593&volume=20&pages=1172-1179&publication_year=2017&author=Chan%2CKY)
- Challis, R. C. et al. Adeno-associated virus toolkit to target diverse brain cells. Annu. Rev. Neurosci. 45, 447–469 (2022).
[Article](https://doi.org/10.1146%2Fannurev-neuro-111020-100834) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhtVChu77L) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35440143) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Adeno-associated%20virus%20toolkit%20to%20target%20diverse%20brain%20cells&journal=Annu.%20Rev.%20Neurosci.&doi=10.1146%2Fannurev-neuro-111020-100834&volume=45&pages=447-469&publication_year=2022&author=Challis%2CRC)
- Byrne, L. C. et al. In vivo-directed evolution of adeno-associated virus in the primate retina. JCI Insight 5, e135112 (2020).
[Article](https://doi.org/10.1172%2Fjci.insight.135112) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32271719) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7259523) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=In%20vivo-directed%20evolution%20of%20adeno-associated%20virus%20in%20the%20primate%20retina&journal=JCI%20Insight&doi=10.1172%2Fjci.insight.135112&volume=5&publication_year=2020&author=Byrne%2CLC)
- Hendriks, S. et al. Ethical challenges of risk, informed consent, and posttrial responsibilities in human research with neural devices: a review. JAMA Neurol. 76, 1506–1514 (2019).
[Article](https://doi.org/10.1001%2Fjamaneurol.2019.3523) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31621797) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9395156) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Ethical%20challenges%20of%20risk%2C%20informed%20consent%2C%20and%20posttrial%20responsibilities%20in%20human%20research%20with%20neural%20devices%3A%20a%20review&journal=JAMA%20Neurol.&doi=10.1001%2Fjamaneurol.2019.3523&volume=76&pages=1506-1514&publication_year=2019&author=Hendriks%2CS)
- Loyola-Gonzalez, O. Black-box vs. white-box: understanding their advantages and weaknesses from a practical point of view. IEEE Access 7, 154096–154113 (2019).
[Article](https://doi.org/10.1109%2FACCESS.2019.2949286) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Black-box%20vs.%20white-box%3A%20understanding%20their%20advantages%20and%20weaknesses%20from%20a%20practical%20point%20of%20view&journal=IEEE%20Access&doi=10.1109%2FACCESS.2019.2949286&volume=7&pages=154096-154113&publication_year=2019&author=Loyola-Gonzalez%2CO)
- McLean, S. et al. The risks associated with artificial general intelligence: a systematic review. J. Exp. Theor. Artif. Intell. 35, 649–663 (2023).
[Article](https://doi.org/10.1080%2F0952813X.2021.1964003) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20risks%20associated%20with%20artificial%20general%20intelligence%3A%20a%20systematic%20review&journal=J.%20Exp.%20Theor.%20Artif.%20Intell.&doi=10.1080%2F0952813X.2021.1964003&volume=35&pages=649-663&publication_year=2023&author=McLean%2CS)
- Feinsinger, A. et al. Ethical commitments, principles, and practices guiding intracranial neuroscientific research in humans. Neuron 110, 188–194 (2022).
[Article](https://doi.org/10.1016%2Fj.neuron.2021.11.011) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhsFCrurc%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35051364) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9417025) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Ethical%20commitments%2C%20principles%2C%20and%20practices%20guiding%20intracranial%20neuroscientific%20research%20in%20humans&journal=Neuron&doi=10.1016%2Fj.neuron.2021.11.011&volume=110&pages=188-194&publication_year=2022&author=Feinsinger%2CA)
- Kaiser, J. & Brainard, J. Ready, set, share. Science 379, 322–325 (2023).
[Article](https://doi.org/10.1126%2Fscience.adg8142) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2023Sci...379..322K) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36701457) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Ready%2C%20set%2C%20share&journal=Science&doi=10.1126%2Fscience.adg8142&volume=379&pages=322-325&publication_year=2023&author=Kaiser%2CJ&author=Brainard%2CJ)
- von Thenen, N., Ayday, E. & Cicek, A. E. Re-identification of individuals in genomic data-sharing beacons via allele inference. Bioinformatics 35, 365–371 (2019).
[Article](https://doi.org/10.1093%2Fbioinformatics%2Fbty643) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Re-identification%20of%20individuals%20in%20genomic%20data-sharing%20beacons%20via%20allele%20inference&journal=Bioinformatics&doi=10.1093%2Fbioinformatics%2Fbty643&volume=35&pages=365-371&publication_year=2019&author=Thenen%2CN&author=Ayday%2CE&author=Cicek%2CAE)
- Jansen, P. R. et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat. Genet. 51, 394–403 (2019).
[Article](https://doi.org/10.1038%2Fs41588-018-0333-3) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXmsVyqs78%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=30804565) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Genome-wide%20analysis%20of%20insomnia%20in%201%2C331%2C010%20individuals%20identifies%20new%20risk%20loci%20and%20functional%20pathways&journal=Nat.%20Genet.&doi=10.1038%2Fs41588-018-0333-3&volume=51&pages=394-403&publication_year=2019&author=Jansen%2CPR)
- Choudhury, S., Fishman, J. R., McGowan, M. L. & Juengst, E. T. Big data, open science and the brain: lessons learned from genomics. Front. Hum. Neurosci. 8, 239 (2014).
[Article](https://doi.org/10.3389%2Ffnhum.2014.00239) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=24904347) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4032989) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Big%20data%2C%20open%20science%20and%20the%20brain%3A%20lessons%20learned%20from%20genomics&journal=Front.%20Hum.%20Neurosci.&doi=10.3389%2Ffnhum.2014.00239&volume=8&publication_year=2014&author=Choudhury%2CS&author=Fishman%2CJR&author=McGowan%2CML&author=Juengst%2CET)
- Van Horn, J. D. & Ball, C. A. Domain-specific data sharing in neuroscience: what do we have to learn from each other. Neuroinformatics 6, 117–121 (2008).
[Article](https://link.springer.com/doi/10.1007/s12021-008-9019-9) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=18473189) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3319052) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Domain-specific%20data%20sharing%20in%20neuroscience%3A%20what%20do%20we%20have%20to%20learn%20from%20each%20other&journal=Neuroinformatics&doi=10.1007%2Fs12021-008-9019-9&volume=6&pages=117-121&publication_year=2008&author=Horn%2CJD&author=Ball%2CCA)
- Ford, D. et al. Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. Am. J. Hum. Genet. 62, 676–689 (1998).
[Article](https://doi.org/10.1086%2F301749) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DyaK1MXlvFeh) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=9497246) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1376944) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Genetic%20heterogeneity%20and%20penetrance%20analysis%20of%20the%20BRCA1%20and%20BRCA2%20genes%20in%20breast%20cancer%20families&journal=Am.%20J.%20Hum.%20Genet.&doi=10.1086%2F301749&volume=62&pages=676-689&publication_year=1998&author=Ford%2CD)
- Maury, E. A. et al. Schizophrenia-associated somatic copy-number variants from 12,834 cases reveal recurrent NRXN1 and ABCB11 disruptions. Cell Genomics 3, 100356 (2023).
[Article](https://doi.org/10.1016%2Fj.xgen.2023.100356) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXhsVWrtL3K) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37601975) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10435376) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Schizophrenia-associated%20somatic%20copy-number%20variants%20from%2012%2C834%20cases%20reveal%20recurrent%20NRXN1%20and%20ABCB11%20disruptions&journal=Cell%20Genomics&doi=10.1016%2Fj.xgen.2023.100356&volume=3&publication_year=2023&author=Maury%2CEA)
- Paré, G. et al. Effects of CYP2C19 genotype on outcomes of clopidogrel treatment. N. Engl. J. Med. 363, 1704–1714 (2010).
[Article](https://doi.org/10.1056%2FNEJMoa1008410) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=20979470) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Effects%20of%20CYP2C19%20genotype%20on%20outcomes%20of%20clopidogrel%20treatment&journal=N.%20Engl.%20J.%20Med.&doi=10.1056%2FNEJMoa1008410&volume=363&pages=1704-1714&publication_year=2010&author=Par%C3%A9%2CG)
- Mellinghoff, I. K. et al. Vorasidenib in IDH1- or IDH2-mutant low-grade glioma. N. Engl. J. Med. 389, 589–601 (2023).
[Article](https://doi.org/10.1056%2FNEJMoa2304194) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXhslWlt7zP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37272516) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Vorasidenib%20in%20IDH1-%20or%20IDH2-mutant%20low-grade%20glioma&journal=N.%20Engl.%20J.%20Med.&doi=10.1056%2FNEJMoa2304194&volume=389&pages=589-601&publication_year=2023&author=Mellinghoff%2CIK)
- Wong, C. UK first to approve CRISPR treatment for diseases: what you need to know. Nature 623, 676–677 (2023).
[Article](https://doi.org/10.1038%2Fd41586-023-03590-6) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2023Natur.623..676W) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXisVCgu7bE) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37974039) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=UK%20first%20to%20approve%20CRISPR%20treatment%20for%20diseases%3A%20what%20you%20need%20to%20know&journal=Nature&doi=10.1038%2Fd41586-023-03590-6&volume=623&pages=676-677&publication_year=2023&author=Wong%2CC)
- Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
[Article](https://doi.org/10.1038%2Fs41586-019-1195-2) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2019Natur.570..332M) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXhtVajsrjI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31042697) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6865822) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Single-cell%20transcriptomic%20analysis%20of%20Alzheimer%E2%80%99s%20disease&journal=Nature&doi=10.1038%2Fs41586-019-1195-2&volume=570&pages=332-337&publication_year=2019&author=Mathys%2CH)
- Yang, A. C. et al. A human brain vascular atlas reveals diverse mediators of Alzheimer’s risk. Nature 603, 885–892 (2022).
[Article](https://doi.org/10.1038%2Fs41586-021-04369-3) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2022Natur.603..885Y) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xjs12kurY%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35165441) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9635042) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20human%20brain%20vascular%20atlas%20reveals%20diverse%20mediators%20of%20Alzheimer%E2%80%99s%20risk&journal=Nature&doi=10.1038%2Fs41586-021-04369-3&volume=603&pages=885-892&publication_year=2022&author=Yang%2CAC)
- Cain, A. et al. Multicellular communities are perturbed in the aging human brain and Alzheimer’s disease. Nat. Neurosci. 26, 1267–1280 (2023).
[Article](https://doi.org/10.1038%2Fs41593-023-01356-x) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXht1GkurvK) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37336975) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10789499) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Multicellular%20communities%20are%20perturbed%20in%20the%20aging%20human%20brain%20and%20Alzheimer%E2%80%99s%20disease&journal=Nat.%20Neurosci.&doi=10.1038%2Fs41593-023-01356-x&volume=26&pages=1267-1280&publication_year=2023&author=Cain%2CA)
- Yuste, R. et al. A community-based transcriptomics classification and nomenclature of neocortical cell types. Nat. Neurosci. 23, 1456–1468 (2020).
[Article](https://doi.org/10.1038%2Fs41593-020-0685-8) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXhs1KhtL%2FL) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=32839617) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC7683348) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=A%20community-based%20transcriptomics%20classification%20and%20nomenclature%20of%20neocortical%20cell%20types&journal=Nat.%20Neurosci.&doi=10.1038%2Fs41593-020-0685-8&volume=23&pages=1456-1468&publication_year=2020&author=Yuste%2CR)
- Speir, M. L. et al. UCSC Cell Browser: visualize your single-cell data. Bioinformatics 37, 4578–4580 (2021).
[Article](https://doi.org/10.1093%2Fbioinformatics%2Fbtab503) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhsFGqur0%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34244710) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8652023) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=UCSC%20Cell%20Browser%3A%20visualize%20your%20single-cell%20data&journal=Bioinformatics&doi=10.1093%2Fbioinformatics%2Fbtab503&volume=37&pages=4578-4580&publication_year=2021&author=Speir%2CML)
- Wan, Z. et al. Sociotechnical safeguards for genomic data privacy. Nat. Rev. Genet. 23, 429–445 (2022).
[Article](https://doi.org/10.1038%2Fs41576-022-00455-y) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XmtFOrsr4%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35246669) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8896074) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Sociotechnical%20safeguards%20for%20genomic%20data%20privacy&journal=Nat.%20Rev.%20Genet.&doi=10.1038%2Fs41576-022-00455-y&volume=23&pages=429-445&publication_year=2022&author=Wan%2CZ)
- Ngai, J. BRAIN 2.0: transforming neuroscience. Cell 185, 4–8 (2022).
[Article](https://doi.org/10.1016%2Fj.cell.2021.11.037) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38Xlt1yksw%3D%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34995517) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=BRAIN%202.0%3A%20transforming%20neuroscience&journal=Cell&doi=10.1016%2Fj.cell.2021.11.037&volume=185&pages=4-8&publication_year=2022&author=Ngai%2CJ)
- Reardon, J. et al. Trustworthiness matters: building equitable and ethical science. Cell 186, 894–898 (2023).
[Article](https://doi.org/10.1016%2Fj.cell.2023.01.008) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXit1Oitrc%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36724788) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10580253) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Trustworthiness%20matters%3A%20building%20equitable%20and%20ethical%20science&journal=Cell&doi=10.1016%2Fj.cell.2023.01.008&volume=186&pages=894-898&publication_year=2023&author=Reardon%2CJ)
- Kowal, E. et al. Community partnerships are fundamental to ethical ancient DNA research. HGG Adv. 4, 100161 (2023).
[CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXht1ygtbc%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37101579) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10123407) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Community%20partnerships%20are%20fundamental%20to%20ethical%20ancient%20DNA%20research&journal=HGG%20Adv.&volume=4&publication_year=2023&author=Kowal%2CE)
- Shim, J. K. et al. Community engagement in precision medicine research: organizational practices and their impacts for equity. AJOB Empir. Bioeth. 14, 185–196 (2023).
[Article](https://doi.org/10.1080%2F23294515.2023.2201478) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37126431) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10615663) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Community%20engagement%20in%20precision%20medicine%20research%3A%20organizational%20practices%20and%20their%20impacts%20for%20equity&journal=AJOB%20Empir.%20Bioeth.&doi=10.1080%2F23294515.2023.2201478&volume=14&pages=185-196&publication_year=2023&author=Shim%2CJK)
- Han, H. R. et al. Exploring community engaged research experiences and preferences: a multi-level qualitative investigation. Res. Involv. Engagem. 7, 19 (2021).
[Article](https://link.springer.com/doi/10.1186/s40900-021-00261-6) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33785074) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8008581) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Exploring%20community%20engaged%20research%20experiences%20and%20preferences%3A%20a%20multi-level%20qualitative%20investigation&journal=Res.%20Involv.%20Engagem.&doi=10.1186%2Fs40900-021-00261-6&volume=7&publication_year=2021&author=Han%2CHR)
- Vollger, M. R. et al. Segmental duplications and their variation in a complete human genome. Science 376, eabj6965 (2022).
[Article](https://doi.org/10.1126%2Fscience.abj6965) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XpvVyns7Y%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35357917) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8979283) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Segmental%20duplications%20and%20their%20variation%20in%20a%20complete%20human%20genome&journal=Science&doi=10.1126%2Fscience.abj6965&volume=376&publication_year=2022&author=Vollger%2CMR)
- Wang, T. et al. The Human Pangenome Project: a global resource to map genomic diversity. Nature 604, 437–446 (2022).
[Article](https://doi.org/10.1038%2Fs41586-022-04601-8) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2022Natur.604..437W) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XhtVGrsbnI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=35444317) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC9402379) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20Human%20Pangenome%20Project%3A%20a%20global%20resource%20to%20map%20genomic%20diversity&journal=Nature&doi=10.1038%2Fs41586-022-04601-8&volume=604&pages=437-446&publication_year=2022&author=Wang%2CT)
- Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
[Article](https://doi.org/10.1038%2Fs41586-019-1506-7) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2019Natur.573...61H) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC1MXhs1agtbvI) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=31435019) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC6919571) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Conserved%20cell%20types%20with%20divergent%20features%20in%20human%20versus%20mouse%20cortex&journal=Nature&doi=10.1038%2Fs41586-019-1506-7&volume=573&pages=61-68&publication_year=2019&author=Hodge%2CRD)
- Berg, J. et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature 598, 151–158 (2021).
[Article](https://doi.org/10.1038%2Fs41586-021-03813-8) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2021Natur.598..151B) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3MXit1WqsbnF) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=34616067) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8494638) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Human%20neocortical%20expansion%20involves%20glutamatergic%20neuron%20diversification&journal=Nature&doi=10.1038%2Fs41586-021-03813-8&volume=598&pages=151-158&publication_year=2021&author=Berg%2CJ)
- Jorstad, N. L. et al. Transcriptomic cytoarchitecture reveals principles of human neocortex organization. Science 382, eadf6812 (2023).
[Article](https://doi.org/10.1126%2Fscience.adf6812) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXitFygtbfP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=37824655) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Transcriptomic%20cytoarchitecture%20reveals%20principles%20of%20human%20neocortex%20organization&journal=Science&doi=10.1126%2Fscience.adf6812&volume=382&publication_year=2023&author=Jorstad%2CNL)
- Elston, G. N. Cortex, cognition and the cell: new insights into the pyramidal neuron and prefrontal function. Cereb. Cortex 13, 1124–1138 (2003).
[Article](https://doi.org/10.1093%2Fcercor%2Fbhg093) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=14576205) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Cortex%2C%20cognition%20and%20the%20cell%3A%20new%20insights%20into%20the%20pyramidal%20neuron%20and%20prefrontal%20function&journal=Cereb.%20Cortex&doi=10.1093%2Fcercor%2Fbhg093&volume=13&pages=1124-1138&publication_year=2003&author=Elston%2CGN)
- Hill, R. S. & Walsh, C. A. Molecular insights into human brain evolution. Nature 437, 64–67 (2005).
[Article](https://doi.org/10.1038%2Fnature04103) [ADS](http://adsabs.harvard.edu/cgi-bin/nph-data_query?link_type=ABSTRACT&bibcode=2005Natur.437...64H) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BD2MXpslaltr0%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=16136130) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Molecular%20insights%20into%20human%20brain%20evolution&journal=Nature&doi=10.1038%2Fnature04103&volume=437&pages=64-67&publication_year=2005&author=Hill%2CRS&author=Walsh%2CCA)
- Markov, N. T. et al. Cortical high-density counterstream architectures. Science 342, 1238406 (2013).
[Article](https://doi.org/10.1126%2Fscience.1238406) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=24179228) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3905047) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Cortical%20high-density%20counterstream%20architectures&journal=Science&doi=10.1126%2Fscience.1238406&volume=342&publication_year=2013&author=Markov%2CNT)
- Nowakowski, T. J., Pollen, A. A., Sandoval-Espinosa, C. & Kriegstein, A. R. Transformation of the radial glia scaffold demarcates two stages of human cerebral cortex development. Neuron 91, 1219–1227 (2016).
[Article](https://doi.org/10.1016%2Fj.neuron.2016.09.005) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BC28XhsFGrtbzJ) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=27657449) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC5087333) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Transformation%20of%20the%20radial%20glia%20scaffold%20demarcates%20two%20stages%20of%20human%20cerebral%20cortex%20development&journal=Neuron&doi=10.1016%2Fj.neuron.2016.09.005&volume=91&pages=1219-1227&publication_year=2016&author=Nowakowski%2CTJ&author=Pollen%2CAA&author=Sandoval-Espinosa%2CC&author=Kriegstein%2CAR)
- Huttenlocher, P. R. & Dabholkar, A. S. Regional differences in synaptogenesis in human cerebral cortex. J. Comp. Neurol. 387, 167–178 (1997).
[Article](https://doi.org/10.1002%2F%28SICI%291096-9861%2819971020%29387%3A2%3C167%3A%3AAID-CNE1%3E3.0.CO%3B2-Z) [CAS](https://www.nature.com/articles/cas-redirect/1:STN:280:DyaK2svnsF2ksg%3D%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=9336221) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Regional%20differences%20in%20synaptogenesis%20in%20human%20cerebral%20cortex&journal=J.%20Comp.%20Neurol.&doi=10.1002%2F%28SICI%291096-9861%2819971020%29387%3A2%3C167%3A%3AAID-CNE1%3E3.0.CO%3B2-Z&volume=387&pages=167-178&publication_year=1997&author=Huttenlocher%2CPR&author=Dabholkar%2CAS)
- Vanderhaeghen, P. & Polleux, F. Developmental mechanisms underlying the evolution of human cortical circuits. Nat. Rev. Neurosci. 24, 213–232 (2023).
[Article](https://doi.org/10.1038%2Fs41583-023-00675-z) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3sXjtlSrsLs%3D) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36792753) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC10064077) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Developmental%20mechanisms%20underlying%20the%20evolution%20of%20human%20cortical%20circuits&journal=Nat.%20Rev.%20Neurosci.&doi=10.1038%2Fs41583-023-00675-z&volume=24&pages=213-232&publication_year=2023&author=Vanderhaeghen%2CP&author=Polleux%2CF)
- Roberts, R. J. V., Pop, S. & Prieto-Godino, L. L. Evolution of central neural circuits: state of the art and perspectives. Nat. Rev. Neurosci. 23, 725–743 (2022).
[Article](https://doi.org/10.1038%2Fs41583-022-00644-y) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB38XislCjsb7N) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=36289403) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Evolution%20of%20central%20neural%20circuits%3A%20state%20of%20the%20art%20and%20perspectives&journal=Nat.%20Rev.%20Neurosci.&doi=10.1038%2Fs41583-022-00644-y&volume=23&pages=725-743&publication_year=2022&author=Roberts%2CRJV&author=Pop%2CS&author=Prieto-Godino%2CLL)
- Kebschull, J. M. et al. Cerebellar nuclei evolved by repeatedly duplicating a conserved cell-type set. Science 370, eabd5059 (2020).
[Article](https://doi.org/10.1126%2Fscience.abd5059) [CAS](https://www.nature.com/articles/cas-redirect/1:CAS:528:DC%2BB3cXis1KksL%2FP) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=33335034) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC8510508) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=Cerebellar%20nuclei%20evolved%20by%20repeatedly%20duplicating%20a%20conserved%20cell-type%20set&journal=Science&doi=10.1126%2Fscience.abd5059&volume=370&publication_year=2020&author=Kebschull%2CJM)
- Defelipe, J. The evolution of the brain, the human nature of cortical circuits, and intellectual creativity. Front. Neuroanat. 5, 29 (2011).
[Article](https://doi.org/10.3389%2Ffnana.2011.00029) [PubMed](http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=21647212) [PubMed Central](http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3098448) [Google Scholar](http://scholar.google.com/scholar_lookup?&title=The%20evolution%20of%20the%20brain%2C%20the%20human%20nature%20of%20cortical%20circuits%2C%20and%20intellectual%20creativity&journal=Front.%20Neuroanat.&doi=10.3389%2Ffnana.2011.00029&volume=5&publication_year=2011&author=Defelipe%2CJ)
Acknowledgements
This work was supported by gifts from Fred Ehrsam, William K. Bowes Jr Foundation and Schmidt Futures; grants from the NIH: R25NS070680 (to A.T.L.), DP2-NINDS 1DP2NS122550-01, U01 U01MH130995 and R21 1R21NS123461-01A1 (to M.F.P.); the Klingenstein–Simons Award in Neuroscience; the Sontag Foundation Distinguished Scientist Award; the Broad Foundation Innovation Award to T.J.N.; and the Roberta and Oscar Gregory Endowment in Stroke and Brain Research to M.F.P. T.J.N. is a New York Stem Cell Foundation Robertson Neuroscience Investigator.
Author information
Authors and Affiliations
-
Department of Neurological Surgery, University of California, San Francisco, San Francisco, CA, USA
Anthony T. Lee, Edward F. Chang & Tomasz J. Nowakowski
-
Weill Institute for Neurosciences, University of California, San Francisco, San Francisco, CA, USA
Anthony T. Lee, Edward F. Chang & Tomasz J. Nowakowski
-
Department of Neurology, University of California, San Francisco, San Francisco, CA, USA
Mercedes F. Paredes
-
Department of Anatomy, University of California, San Francisco, San Francisco, CA, USA
Tomasz J. Nowakowski
-
Department of Psychiatry and Behavioral Sciences, University of California, San Francisco, San Francisco, CA, USA
Tomasz J. Nowakowski
-
Eli and Edythe Broad Center for Regeneration Medicine and Stem Cell Research, University of California, San Francisco, San Francisco, CA, USA
Tomasz J. Nowakowski
Contributions
All authors contributed to the writing and review of the final manuscript. E.F.C. and T.J.N. were involved in the initial conception of the paper.
Corresponding author
Correspondence to Tomasz J. Nowakowski.
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Josef Parvizi, Khara Ramos, Ziv Williams and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, A.T., Chang, E.F., Paredes, M.F. et al. Large-scale neurophysiology and single-cell profiling in human neuroscience. Nature 630, 587–595 (2024). https://doi.org/10.1038/s41586-024-07405-0
-
Received06 July 2023
-
Accepted09 April 2024
-
Published19 June 2024
-
Issue Date20 June 2024
-
DOIhttps://doi.org/10.1038/s41586-024-07405-0
Subjects
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.